Automated and Assisted Solid-Phase Extraction (SPE) Sample Preparation for Bioanalysis of GLP-1 Analogs from Plasma
Makda Araya, Chelsea Plummer
Waters Corporation, United States
Published on May 1, 2026
Abstract
The expansion of glucagon-like peptide 1 (GLP-1) analogs, beyond their initial use for type II diabetes mellitus into obesity and neurodegenerative research, necessitates highly sensitive and reproducible bioanalytical methods. These synthetic analogs are modified to resist enzymatic degradation, which increases in vivo duration but complicates pharmacokinetic (PK) studies due to high non-specific binding (NSB) and matrix interference. This work demonstrates a rapid, two-step SPE protocol using Oasis™ PRiME HLB μElution™ Plates for the extraction of five GLP-1 analogs: semaglutide, exenatide, liraglutide, lixisenatide, and tirzepatide from plasma. By incorporating QuanRecovery™ Collection Plates, NSB can be mitigated. The extraction procedure was evaluated using an automated and an assisted, semi-automated protocol with the Andrew+™ and Pipette+™ connected devices. This combined workflow provides a simple, high-throughput solution for the sensitive and accurate quantification of therapeutic peptides in complex biological matrices.
Benefits
- Simple and quick two-step SPE protocol using Oasis PRiME HLB µElution Plate achieved reproducible recoveries of five GLP-1 analogs from rat plasma
- QuanRecovery Collection Plates mitigate NSB, improving recovery
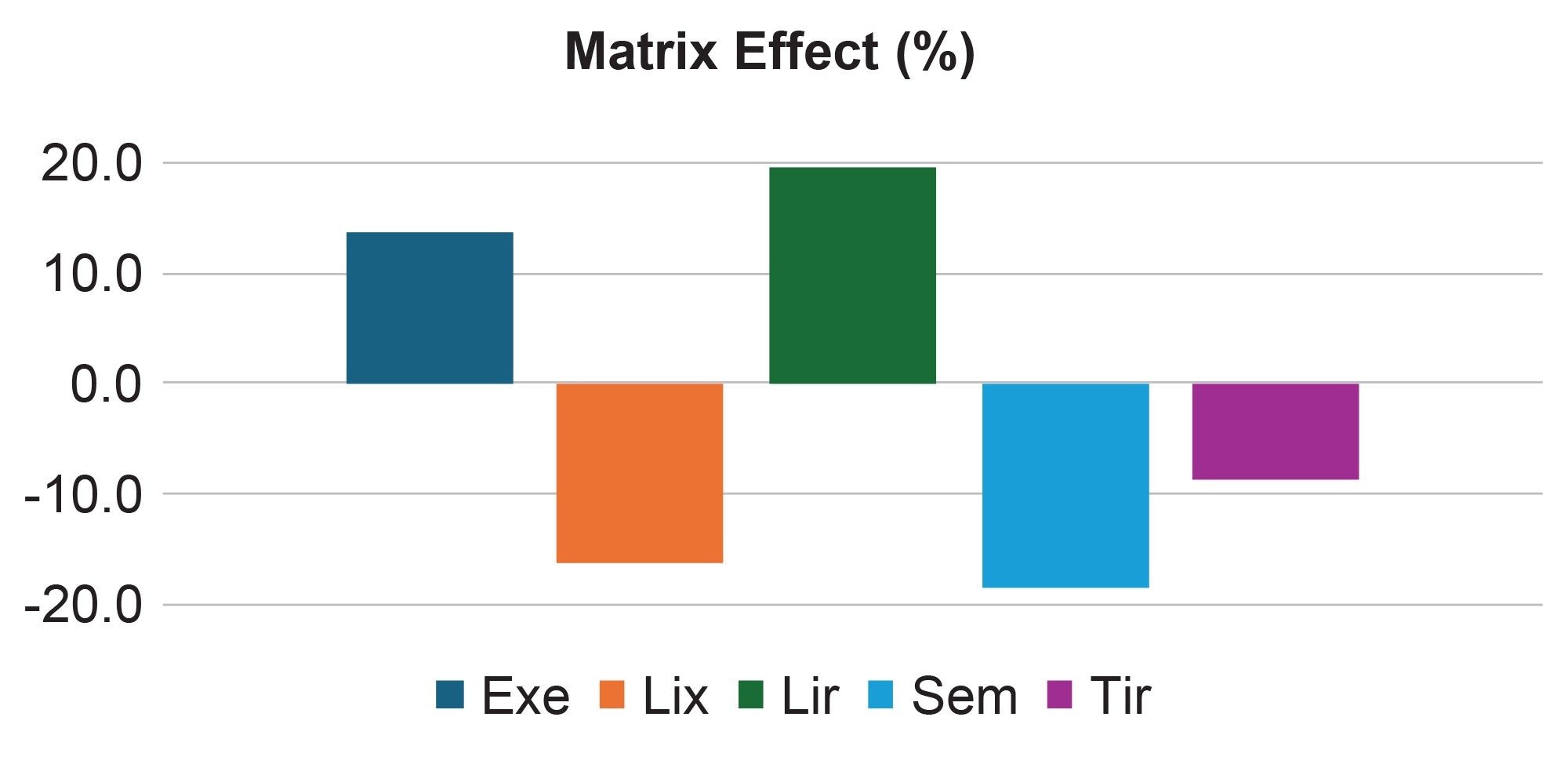
- Low matrix effects (<20%) obtained for all five analytes
- Linear calibration curves obtained in the range of 5-1000 ng/mL for automated and assisted approaches
- Whole protocol executed with Andrew+ Robot fully automated is requires 42% less time than manual execution, and assisted, easy Pipette+ Connected Device requires 54% less time than manual execution
Introduction
GLP-1 analogs, initially intended to treat type II diabetes mellitus, are currently widely used for obesity and cardiovascular health management. In addition, investigation is underway for neurodegenerative and cognitive disorders. As expansions continue into new applications in health, this creates an increased demand for robust, selective, and sensitive bioanalytical methods to assess drug efficacy, safety, immunogenicity and metabolism.
Quantifying these analogs in plasma is inherently difficult because they are modified to resist enzymatic degradation and increase duration of action in vivo. These lipophilic modifications not only lead to significant NSB to plastic and metal surfaces but increase affinity to plasma proteins. In traditional workflows, these sticky peptides are often lost during sample preparation or within the LC fluidics path, leading to poor recovery and decreased sensitivity. Additionally, the presence of co-extracted phospholipids in plasma can cause significant matrix effects, further compromising the accuracy of pharmacokinetic (PK) and metabolism studies.
To address these challenges, a streamlined, broad use, two-step SPE protocol was evaluated. Oasis PRiME HLB, providing high-sensitivity enrichment with sorbent chemistry designed to remove phospholipids, was used in a μElution format, eliminating the need for evaporation and reconstitution steps where peptide loss is most common. While Previous work used Oasis HLB for semaglutide extraction, Oasis PRiME HLB was selected to expand on a general-purpose method for GLP-1s.1 When paired with MaxPeak™ High Performance Surfaces (HPS) technology in the ACQUITY™ Premier System and QuanRecovery plates, this workflow effectively manages surface interactions and NSB. Curve generation, sample pretreatment and SPE steps were fully automated using the Andrew+ Robot in addition to an assisted, semi-automated format with the Pipette+ Smart Pipette Device. Both tools paired with OneLab™ Software provide traceable, audit-trailed and compliant workflows that reduce manual steps by analysts. The result is a fast, reproducible method for the simultaneous quantification of five major GLP-1 analogs, ensuring high data quality without sacrificing laboratory throughput.
Experimental
Exenatide acetate (AA0039QE), liraglutide (AA003R7S), lixisenatide (AA00C6IL), tirzepatide (AA01MVAI) and semaglutide (AA00H208) from aablocks (San Diego, CA) were dissolved in diluted methanol. 200 µL of Sprague Dawley rat plasma from BioIVT (Woodbury, NY) was spiked with the analyte solution before precipitation with 200 µL of cold, acidified acetonitrile. The supernatant was then diluted with 400 µL of acidified water and loaded onto an Oasis PRiME HLB µElution Plate (p/n: 186008052, Milford, MA). After eluting 2x with 25 µL 3% ammonium hydroxide in 75% acetonitrile (Figure 1), eluates were diluted 1:1 with water and directly injected on an ACQUITY Premier System paired with a Waters Xevo™ TQ Absolute Mass Spectrometer. QuanRecovery Collection Plates were used to minimize NSB.

LC Conditions
|
System: |
ACQUITY Premier System with Flow-Through Needle (FTN) |
|
Column: |
ACQUITY Premier Peptide CSH™ C18 Column, 130Å, 1.7 µm, 2.1 x 50 mm (Waters, p/n: 186009487) |
|
Column temperature: |
65 °C |
|
Mobile phase A |
0.1% Formic Acid in 18.2 MΩ*cm Water (v/v) |
|
Mobile phase B: |
0.1% Formic Acid in Acetonitrile (v/v) |
|
Injection volume: |
10 µL |
|
Weak needle wash: |
0.1% Formic Acid in Acetonitrile (v/v) |
|
Strong needle wash: |
10% Acetonitrile in 0.1% Formic Acid (v/v) |
MS Conditions
|
System: |
Xevo TQ Absolute Mass Spectrometer |
|
Ionization mode: |
ESI positive |
|
Capillary voltage: |
2.00 kV |
|
Cone voltage: |
32 V |
|
Desolvation temperature: |
500 °C |
|
Desolvation gas flow: |
1100 L/Hr |
|
Cone gas flow: |
150 L/Hr |
|
MS software: |
MassLynx™ 4.2 Software, TargetLynx™ 4.2 Software |
LC Gradient Table


Automation and Smart Devices
Andrew+ Robot, Pipette+ and Extraction+™ Connected Devices driven by OneLab Software.
Results and Discussion
The extraction of five GLP-1 analogs (semaglutide, exenatide, liraglutide, lixisenatide and tirzepatide) from plasma for LC-MS/MS quantification was evaluated and achieved using three distinct approaches: fully manual, assisted, semi-manual (Pipette+ with Extraction+ Devices), and fully automated (Andrew+ Robot with Extraction+ Device). The procedure was evaluated fully manually to first determine suitability, followed by fully automated and assisted pipetting methods. Depending on the approach used, the total time of the workflow for 48 samples (48 wells) varied. As seen in Figure 2C, a fully manual approach takes around 3 hours 30 minutes, while moving to a fully automated workflow reduces the time to at 2 hours 1 minute. The fully automated approach also greatly reduces the labor time for sample preparation. The assisted Pipette+ method was fastest at 1 hour 36 minutes because a scientist can perform certain steps quicker than the robotic arm. Use of OneLab Software allows all three workflows to be fully traceable for compliance.
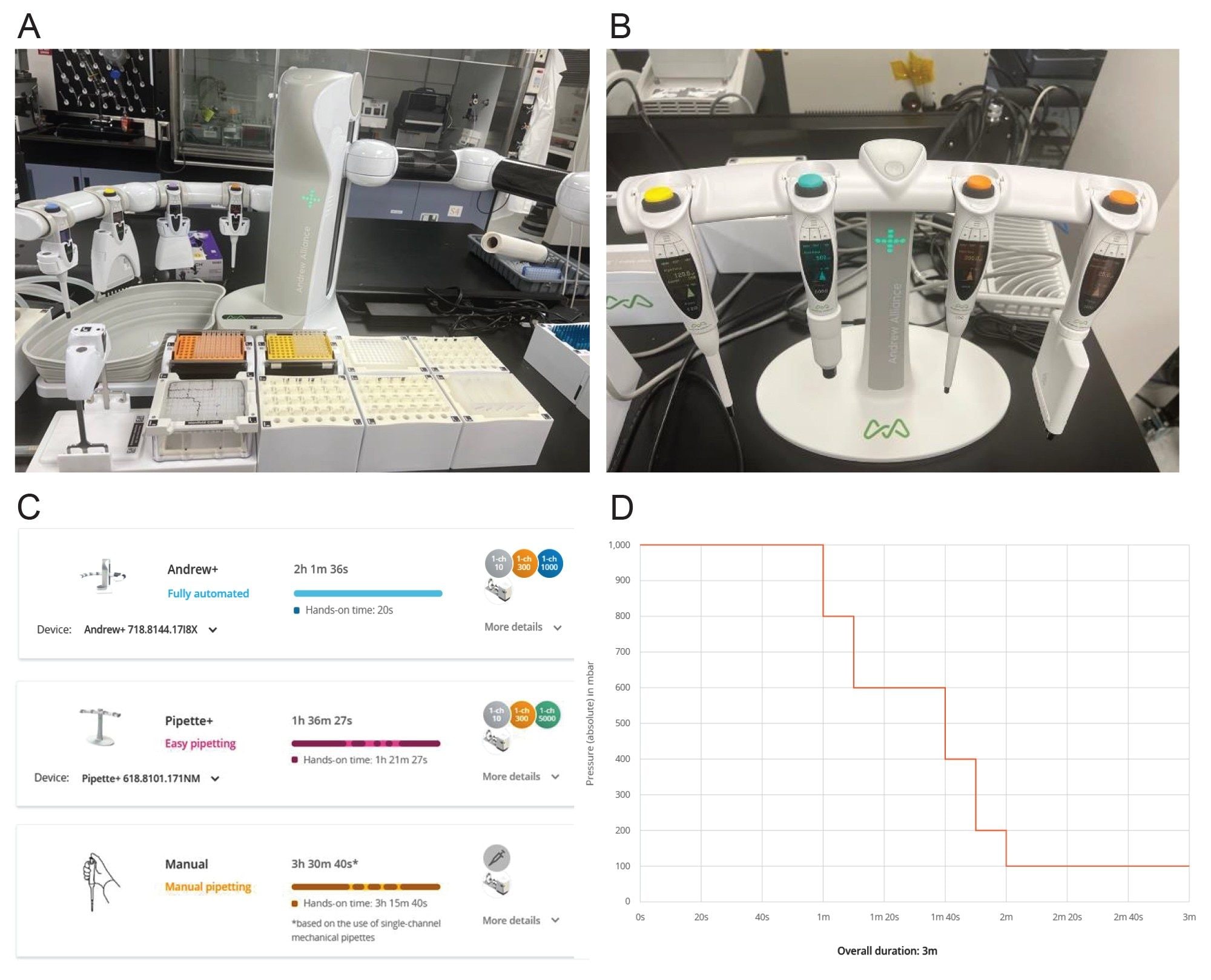
For both workflows, single‑channel automatic pipettes (10 µL, 300 µL, and 1000 µL) and an 8‑channel 120 µL automatic pipette were used. As shown in Figure 2A, the fully automated workflow employs three tip‑insertion, one storage plate, three microtube storage, one deep‑well microplate, and the Extraction+ manifold dominos. In contrast, the Pipette+ workflow only required the Extraction+ manifold domino.

A critical challenge in peptide bioanalysis is balancing recovery with clean extracts. This work required an optimization process to evaluate various protein precipitation (PPT) conditions prior to SPE, as well as evaluating the use of a two- vs three-step SPE protocol for broad use across multiple GLP-1 analogs. Spiked plasma was precipitated with acidified acetonitrile, clarified by centrifugation and then the supernatant was processed using a 2‑step SPE method, balancing cleanliness and speed. Oasis PRiME HLB µElution Plates were chosen for their fast two- and three-step protocols and superior phospholipid cleanup. The µElution format is particularly suited for peptide bioanalysis, as it allows for small volume processing and removes the need for evaporation and reconstitution.
Additionally, the use of QuanRecovery collection plates with high performance surfaces were used to further impact recovery and sensitivity due to minimizing NSB as described previously.2 NSB and surface interactions that contribute to analyte loss were further mitigated by using Protein LoBind tubes for stock storage and sample storage during sample pretreatment steps.
The final optimized method achieved:
- Low matrix effects: Less than 20% for all five GLP-1 analogs (Figure 4)
- Linearity: Calibration curves ranging from 5–1000 ng/mL with R2 > 0.99 across both automated and assisted-pipetting work
As demand for reproducible and reliable data increases, many are turning to automation. Automation offers accurate and reproducible data, higher throughput and faster turnaround, eliminates human bias and error, enhances compliance and traceability and provides efficient workflows that garner long-term cost savings.
This approach shows automated SPE coupled with LC-MS/MS can be successfully applied to quantify diverse GLP-1 peptides in plasma. During the development of the fully automated protocol, a notable challenge was identified involving solvent evaporation during the elution step. In an optimal workflow, temperature needs to be strictly controlled to reduce this risk, as solvent volatility can lead to evaporation and poor reproducibility. Future work to mitigate this risk on the Andrew+ platform could utilize a device with temperature control called Peltier+ Connected Device; however, volumes would need to be scaled to be amenable.
For labs where a Peltier+ is not utilized, the assisted pipetting approach with the Pipette+ device serves as an excellent alternative, reducing the total workflow time by approximately 30 minutes compared to a fully automated approach and minimizing time the elution solvent is exposed to the atmosphere. Both approaches have the optimal benefit of maintaining full electronic records for compliance and review supported by OneLab Software.
Conclusion
The results of this study demonstrate that automated and semi-automated SPE coupled with LC-MS/MS provide a robust and reproducible solution for the quantification of diverse GLP-1 peptides in plasma. By utilizing the Andrew+ Robot and its connected devices, laboratories can offload the manual workflow, leading to a reduction in time of up to 54%, depending on the approach taken, while ensuring full traceability via OneLab software.
References
- Mary Trudeau and Angela Scumaci, SPE-LC/MS Bioanalytical Quantification of the Biotherapeutic Peptide, Semaglutide from Plasma, (2023). Waters Application Note: 720008097.
- Mary Trudeau and Matthew A. Lauber, Improved GLP-1 Receptor Agonist Peptide Recovery Using a QuanRecovery with MaxPeak High Performance Surfaces (HPS) Collection Plate, (2025). Waters Application Note: 720008717.
720009318, May 2026