In this application note we present optimized size-exclusion chromatography (SEC) methods that can separate soluble AAV selfassociated forms and fragments under non-denaturing conditions for several CMV-GFP control AAV serotypes including AAV1, AAV2, AAV5, AAV6, AAV8, and AAV9.
As the development of gene therapy products accelerates, the need to develop sound and efficient analytical strategies to help guide the development of manufacturing processes and evaluate the quality of clinical adeno-associated virus (AAV) vectors has become more important. Among other critical quality attributes, the levels of potential AAV aggregates represented as high molecular weight species (HMWS) and AAV fragments represented as low molecular weight species (LMWS) may also require monitoring.1 Here we present optimized size-exclusion chromatography (SEC) methods that can separate soluble AAV selfassociated forms and fragments under non-denaturing conditions for several CMV-GFP control AAV serotypes including AAV1, AAV2, AAV5, AAV6, AAV8, and AAV9.2

Bulk AAV8-Null and AAV8-GFP control samples were supplied by BioReliance (Rockville, MD). The assorted AAV serotype samples were purchased from Vigene Biosciences (Rockville, MD) and were formulated in PBS. BEH450 SEC Protein Standard Mix resuspended in 500 μL PBS containing 10 μg/mL of L-tryptophan (Sigma).
|
LC conditions |
|
|---|---|
|
Systems: |
ACQUITY UPLC H-Class Bio |
|
Detection: |
ACQUITY UPLC with FLR Detector (analytical flow cell), unless otherwise noted. Wavelengths: excitation at 280 nm and emission at 350 nm Wyatt microDAWN MALS detector (Santa Barbara, CA) Waters ACQUITY RI Detector |
|
Column: |
XBridge Protein BEH SEC, 450 Å, 3.5 μm, 7.8 mm × 300 mm with included BEH450 SEC Protein Standard Mix (p/n: 176003599) |
|
Column temp.: |
25 °C |
|
Sample temp.: |
6 °C |
|
Mobile phase: |
A: 10 mM NaH2PO4, 10 mM NaH2PO4, pH 6.6 (HCl); B: 10 mM NaH2PO4, 10 mM Na2HPO4, 500 mM KCl, pH 6.6 (HCl); All 0.1 μm sterile filtered, unless otherwise noted |
|
Flow rate: |
0.6 mL/min, unless otherwise noted |
|
Sample vials: |
Polypropylene 12 × 32 mm Screw Neck Vial, with Cap and PTFE/silicone Septum, 300 μL Volume (p/n:186002640) |
|
Chromatography software: |
Empower 3 |
In developing an SEC separation for AAV, it is important to recognize the upper analyte size limitation of SEC, which is below the sub-visible range with a detectable upper particle size diameter limit of approximately 100 nm. As aggregate size exceeds 100 nm, it may be disrupted under SEC conditions or trapped by the frits or packed bed of the column. Therefore, the sizing of entities larger than 100 nm will typically fall in the realm of complementary methods such as dynamic light scattering and nanoparticle tracking analysis (NTA), among others.3,4 It should also be noted that significant increases in the levels of subvisible aggregate forms may not be concomitant with an increase in lower valency aggregates such as dimer and trimer observed by SEC profiles.
With these considerations, the SEC separation of AAV monomer, dimer, and lower valency multimers in addition to low molecular weight forms was evaluated on a 450 Å (45 nm) average pore-size BEH diol-bonded SEC column. An SEC particle with an average pore size of 450 Å would not generally be considered adequate for the SEC separation of an analyte with a protein molecular weight of nearly 4000 KDa and a ssDNA molecular weight of approximately 1500 KDa. However, due to the compact structure of the AAV with a diameter of 25 nm, it was predicted that this high efficiency SEC particle with a large pore volume could provide the needed separation with the larger than average size pores providing most of the accessible pore volume for the HMWS. Additionally, while initial method development was undertaken on a 4.6 mm I.D. column packed with 2.5 μm diameter particles, a larger 3.5 μm particle size, 7.8 mm I.D. column was selected for the final method to minimize potential sample sieving effects.
Intrinsic protein fluorescence detection was also employed to provide maximum sensitivity given the low concentrations of many AAV formulated samples and the small injection volumes desired for SEC due to the low product yields of the manufacturing processes being developed. Intrinsic protein fluorescence also has an added advantage that the response factor is not as greatly impacted by changes in the DNA content of the AAV in comparison to UV absorbance.5
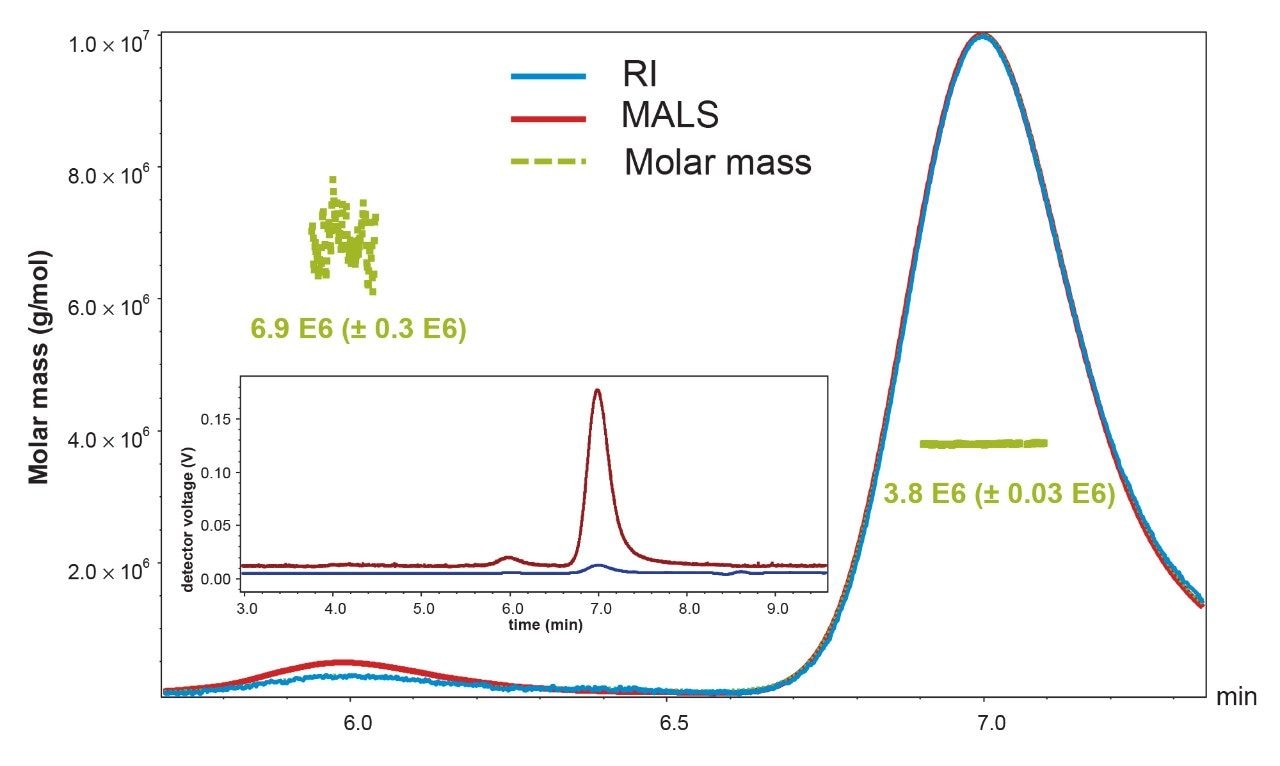
Method development initially used an AAV8-Null control sample due to availability. A null control sample is an AAV capsid that does not contain DNA. The separation observed using a phosphate buffered saline mobile phase (PBS, 10 mM sodium phosphate with 150 mM NaCl) was evaluated using a Wyatt microDAWN Multi-Angle Light Scattering detector (MALS) and an ACQUITY RI (refractive index) Detector (Figure 1). The SEC-MALS data confirmed that separation between the dimeric and monomeric AAV forms was observed. Additionally, putative multimeric forms preceding dimer were also observed by FLR but could not be assigned molecular weights by MALS due to their low abundance.
A single-factor optimization scheme for the mobile phase starting with PBS was employed to maximize aggregate recovery and decrease peak tailing of the monomeric capsid. Wherein, the evaluation of pH while holding ionic strength constant (150 mM NaCl), followed by evaluating ionic strength at the observed optimal pH was accomplished using Auto•Blend Plus Technology.6 Finally, salt types (NaCl, KCl, and sodium perchlorate) and other additives (arginine and isopropanol) were evaluated. Ultimately, a mobile phase consisting of 20 mM sodium phosphate, pH 6.6, with 150 mM KCl was found to provide a functional separation for the AAV8-Null control sample. This single-factor optimization approach was opted for in lieu of a more rigorous full-factorial method development to conserve sample. However, it should be noted that using a full-factorial optimization is generally preferred to determine optimal method conditions since the use of a single-factor optimization scheme may result in settling on a local optimum. In deploying this method for the analysis of DNA containing AAV serotypes (CMV-GFP), it was observed that the optimal concentration of KCl required to maximize the recovery of dimer and multimer varied by serotype. The finalized method conditions and chromatographic profiles for serotypes AAV1, AAV2, AAV5, AAV6, AAV8, and AAV9 are presented in Figure 2.
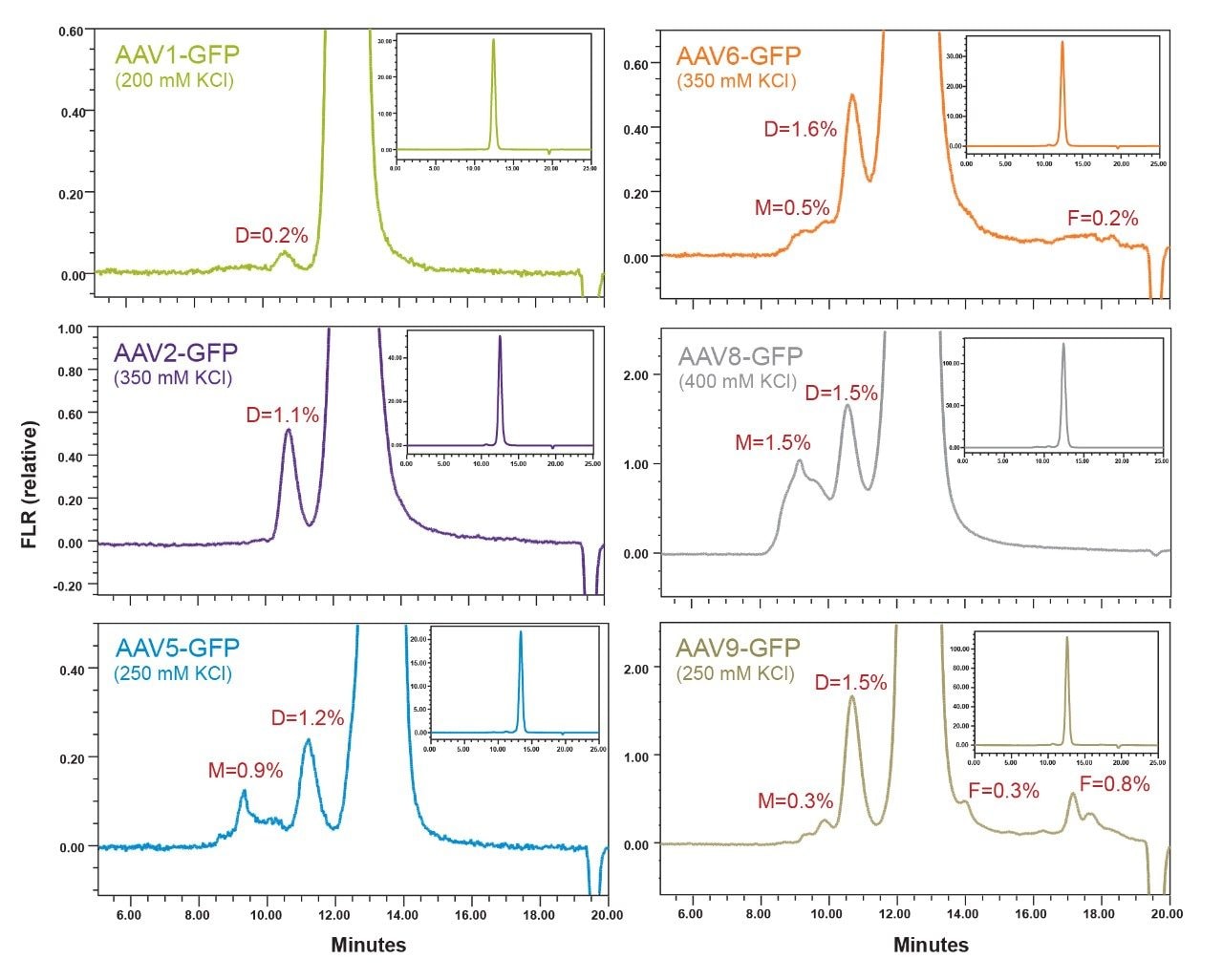
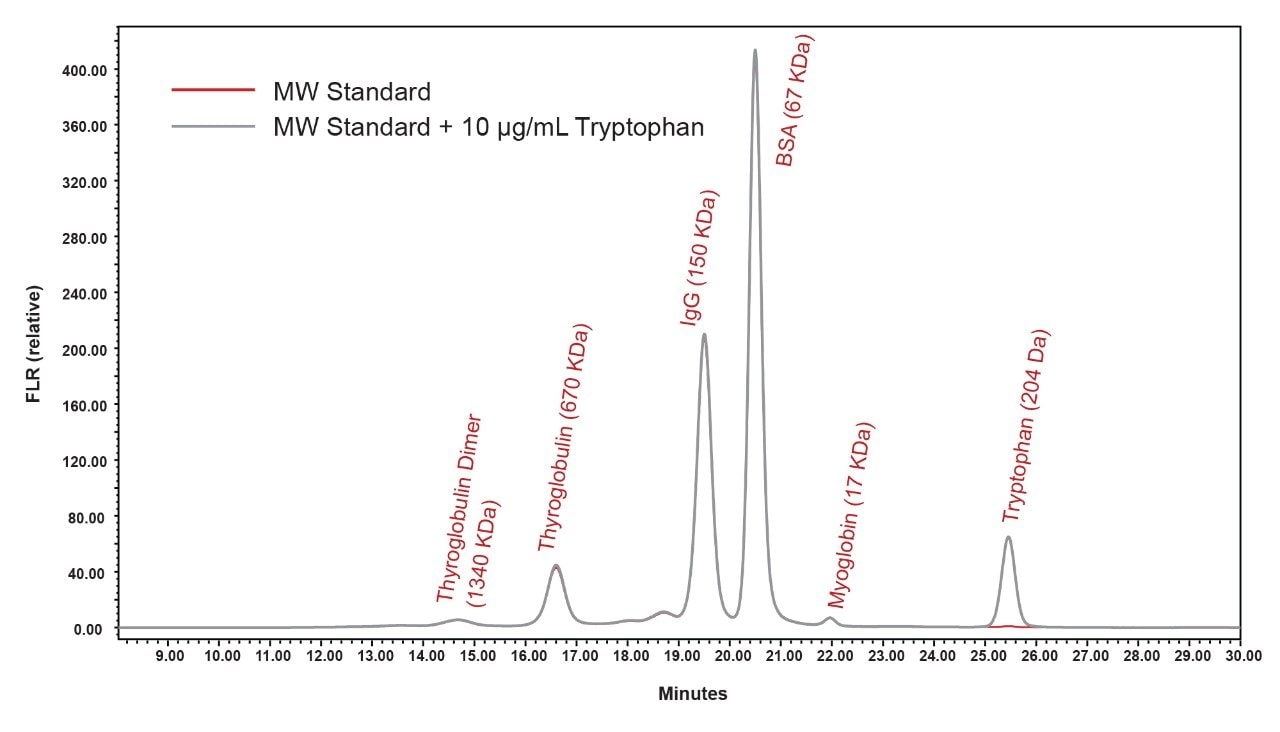
We observe that the peak shapes for both the dimer HMWS and monomeric AAV forms are symmetrical and return to baseline appropriately. In addition, detectable levels of HMWS with retention times that are consistent with multimeric AAV forms were observed for several of the serotypes. Significant amounts of LMW forms were only observed in the AAV9 serotype and to a much lower extent in AAV6. As previously noted, intrinsic protein fluorescence detection was used for this method to enhance the sensitivity for low abundance high molecular weight and low molecular weight forms. However, when evaluating the SEC protein standard mix, the small molecule total permeation volume marker (uracil) is not observed due to low quantum yield. Being able to track the total permeation volume of an SEC column is valuable in SEC method development since in this case any FLR signal persisting after the total permeation volume is indicative of protein that is being excessively retained on the column. Therefore, it is recommended that L-tryptophan at 10 μg/mL be added to the standard to mark the total permeation volume since this amino acid is the primary fluorophore of the protein that is detected when using intrinsic protein fluorescence (Figure 3).
A BEH SEC column with an average pore size of 450 Å and a particle diameter of 3.5 μm was demonstrated to be effective in the separation of AAV monomers from their HMW dimers, lower valency multimers, and LMW fragments. The minimal amount of ionic strength (KCl) required for optimal peak shape and recovery varied by serotype and these levels are reported for serotypes AAV1, AAV2, AAV5, AAV6, AAV8, and AAV9. While a 450 Å average pore size may have been considered less than ideal for this separation, the results demonstrate that the high pore volume and efficiencies of these particles enable an effective separation. It is proposed that the AAV HMWS separation is driven primarily through the distribution of pore sizes larger than 450 Å. Additionally, intrinsic protein fluorescence was used to extend the sensitivity of the method for the AAV samples tested.
The authors would like to thank BioReliance for providing the AAV-8 null and AAV-8 GFP samples.
720006812, March 2020