The BioAccord System controlled by compliant-ready UNIFI Software is purposefully designed with integrated methods for routine therapeutic protein analysis. It can perform routine peptide characterization and accurate mass screening for CQA monitoring. In this application note, we demonstrate peptide mapping and monitoring capabilities of the BioAccord System using forced degradation study of a monoclonal antibody (mAb) as a case study.
Biotherapeutics undergo rigorous characterization and monitoring during their development and manufacturing to establish product quality and safety for regulatory compliance.1,2 High-performance LC-MS technology, often utilized for the characterization of a protein’s chemical and post-translational modifications at the peptide level, has now been expanded beyond the conventional realm to attribute monitoring across different stages of the product life cycle. Routine monitoring of these quality attributes requires fit-for-purpose analytical platforms that are robust, easy to operate, and readily deployable in laboratories across an organization. The BioAccord System is a new, compact, high-performance, LC-MS system purposefully designed to meet such requirements. It consists of an ACQUITY UPLC I-Class PLUS, ACQUITY RDa Detector featuring SmartMS Technology, compliance-ready UNIFI Software, and consumables all tailored to assist biopharmaceutical applications. The UNIFI Scientific Information System provides integrated, analytical workflow methods with automated data acquisition, processing and reporting capabilities allowing peptide monitoring assays to be readily performed and documented for a single or large batch of samples. In this application note, we demonstrate peptide mapping and monitoring capabilities of the BioAccord System using forced degradation3,4 study of a monoclonal antibody (mAb) as a case study.
Heat stress: An aliquot of NISTmAb reference material (NIST RM 8671, Gaithersburg, USA) heated at 37 °C for one week.
Oxidative stress: Two aliquots of NISTmAb reference material was incubated with 0.01% and 3% H2O2 respectively at room temperature for 24 h.
Peptide samples: Intact mAb samples were reduced and alkylated with 0.5 M DTT and iodoacetamide solutions before buffer exchanging into 0.1 M tris (pH 7.6) using NAP- 5 size-exclusion columns (GE Healthcare Life Sciences, Pittsburgh, USA). The samples were digested with trypsin (Promega, Madison, USA) for 4 h at 37 °C (20:1 protein to enzyme) followed by acidification with 10% formic acid.
|
System: |
ACQUITY UPLC I-Class PLUS |
|
Detection: |
ACQUITY TUV |
|
Vials: |
LCGC Certified Clear Glass 12 × 32 mm Screw Neck Total Recovery Vial, with Cap and Preslit PTFE/Silicone Septa, 1 mL (p/n: 186000385C) |
|
Column: |
ACQUITY UPLC CSH C18, 130 Å, 1.7 μm, 1 × 100 mm (p/n: 186006937) |
|
Column temp.: |
60 °C |
|
Sample temp.: |
6 °C |
|
Injection volume: |
5 μL |
|
Flow rate: |
0.2 mL/min |
|
Mobile phase A: |
0.1% formic acid in H2O |
|
Mobile phase B: |
0.1% formic acid in acetonitrile |
|
Gradient: |
1% B hold for 1 min, 1–35% B over 50 min, 35–85% B over 6 min, 85% B hold for 4 min, 85–1% over 6 min and 1% B re-eqilibration for 13 min |
|
System: |
ACQUITY RDa Detector |
|
Ionization mode: |
ESI positive |
|
Acquisition mode: |
Full MS scans with CID fragmentation |
|
Acquisition range: |
m/z 50–2000 |
|
Capillary voltage: |
1.2 kV |
|
Collision energy: |
60–120 V (low/high energy ramping) |
|
Cone voltage: |
30 V |
|
Desolvation energy: |
350 °C |
|
Intelligent data capture: |
On |
|
Informatics software: |
UNIFI Scientific Information System v. 1.9.4 |
This application note highlights the utilization of the BioAccord System for routine peptide quality attribute profiling and monitoring. This includes: identification of peptides and their modifications using peptide mapping data, archiving a selected list of potential quality attributes in a secured user-defined custom library using UNIFI scientific library feature followed by targeted peptide monitoring (Figure 1). During routine monitoring of peptide attributes, the custom library items are retrieved and utilized in the accurate mass screening method for targeted CQA monitoring and relative quantification. To enhance usability, the accurate mass screening method can be programmed to notify the user of any attributes exhibiting unusual modification levels through a thresholding feature (the limit check). Both peptide mapping and accurate mass screening described below are compliance-ready methods integrated in the same informatics platform with the library feature to provide the user with a continuous workflow from peptide attribute profiling to monitoring. The overall analysis is further elaborated using trastuzumab test case data.
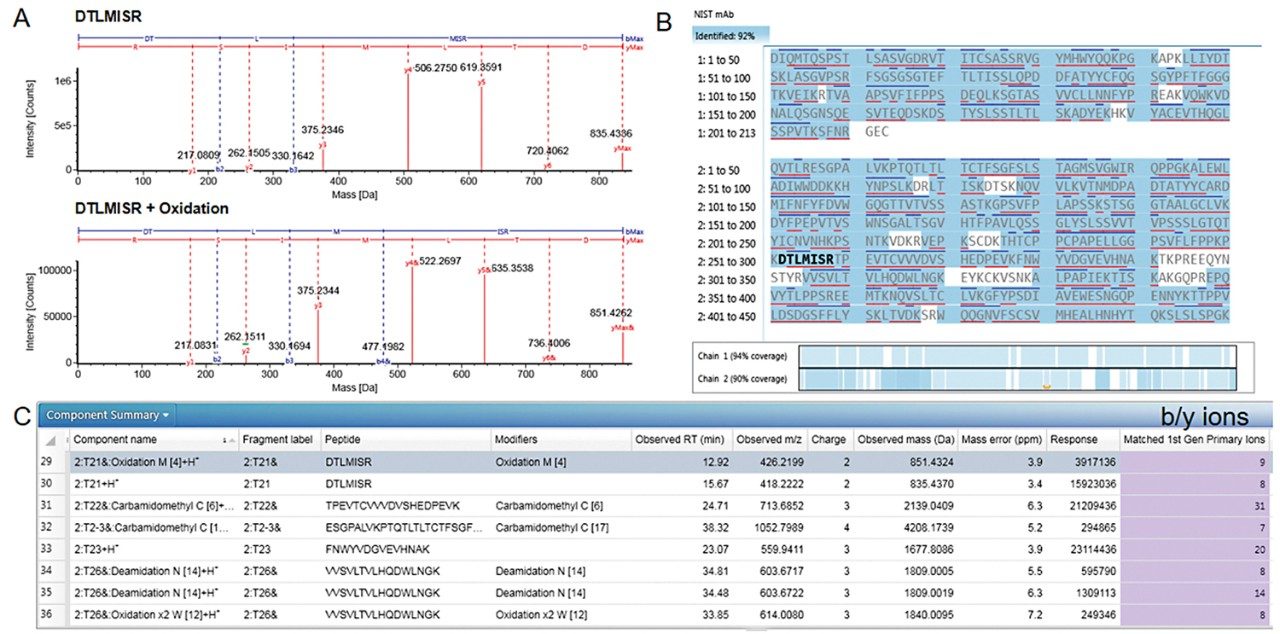
In brief, the mAb sample aliquots were subjected to elevated temperature and chemical oxidation followed by tryptic digestion as stated in the experimental section. The data acquisition was performed using the BioAccord System with alternative MS scans, first with no collision energy and then at elevated collision energy to generate informative fragment ions in data independent acquisition (DIA) mode using an energy ramp from 60–120 V. The acquired data were processed in the peptide mapping method to confirm peptides and their modifications based on the accurate mass, and each assignment was further verified using primary fragment ions for high confidence peptide attribute profiling. The annotated fragmentation spectra for unmodified and modified HC:T21 (Heavy Chain, tryptic peptide number 21 from N-terminus) are shown in Figure 2A. The sequence coverage in Figure 2B includes all peptide sequences verified with ±10 ppm mass accuracy and containing a minimum of 3 b/y fragment ions per peptide. The control and stressed NISTmAb samples reported over 90% sequence coverage. The component table (Figure 2C) provides a comprehensive summary of all verified peptides and modifications identified in each sample along with their respective m/z, retention time, mass tolerance, charge states and number of primary fragment ions (b/y ions). The oxidative stress conditions used for NISTmAb, induced oxidation of both methionine and tryptophan. For routine CQA monitoring, a selected list of peptides from the component table was sent to a custom library. Similarly, the UNIFI processing method described here can be applied to data acquired on other Waters HRMS systems for peptide mapping and PTM confirmation.
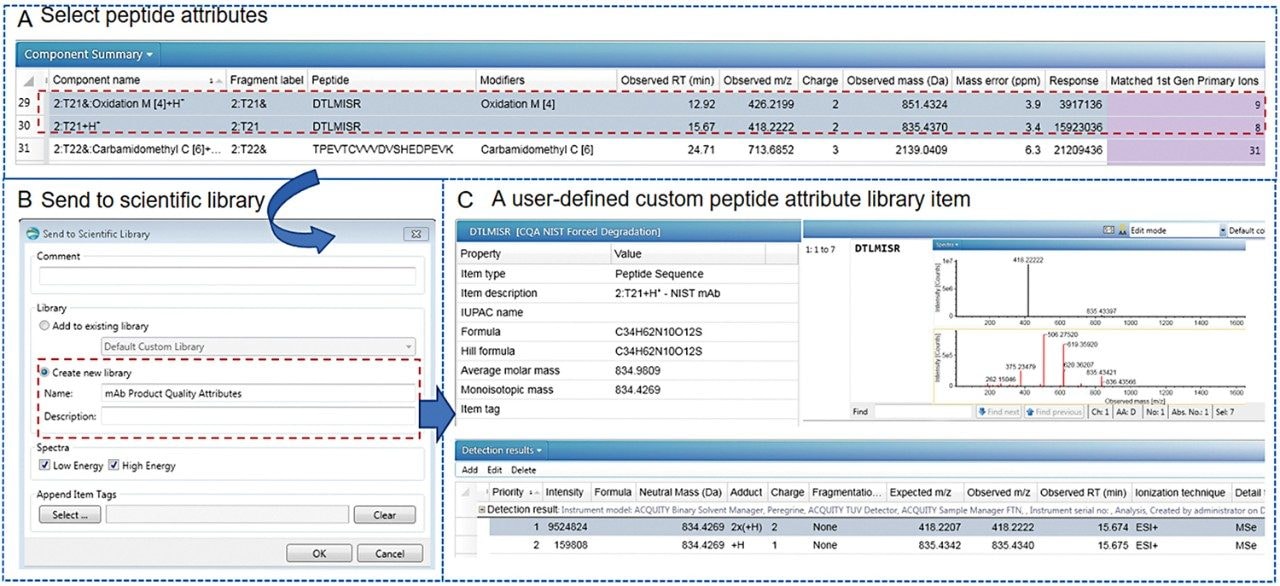
Following peptide identification, a custom library was generated from the peptide mapping data in Step 1. This includes selecting peptide attributes of interest directly from the component table (Figure 3A) and creating a new custom library using “send to library” option in UNIFI (Figure 3B). Each peptide entry consists of several key parameters in addition to peptide sequence, chemical composition and neutral mass such as: retention time, m/z, MS intensity, MS and fragmentation spectra, and charge state information (Figure 3C) that can be selectively retrieved and used in targeted peptide monitoring methods. The case study stated here archived 28 potential peptide quality attributes associated with stressed NISTmAb samples in “mAb Product Quality Attributes” library using UNIFI scientific library feature. These custom libraries can be modified, saved, and shared among other UNIFI users within an organization.
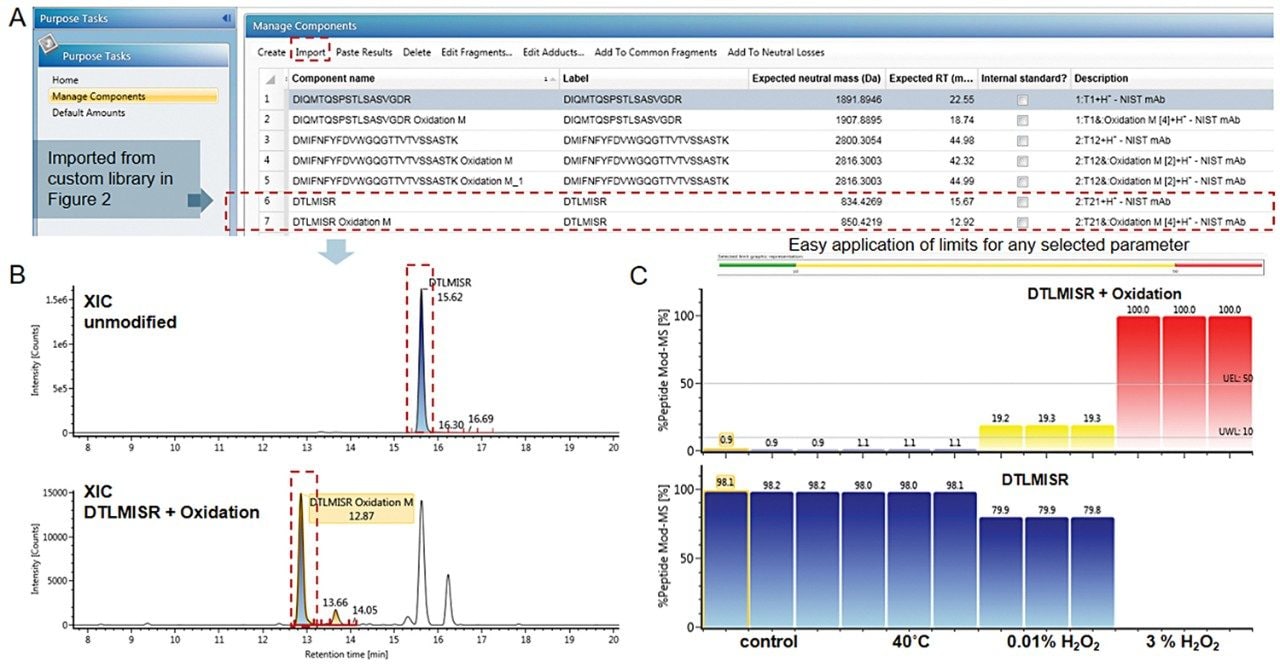
The application note further illustrates the use of UNIFI Accurate Mass Screening data processing method for routine peptide monitoring. With generic processing parameters, intensity thresholding for peptide attributes limit check criteria and reporting; the screening method has made routine peptide monitoring streamlined for improved robustness and productivity. In this case study, the 28-target peptide attributes selected for monitoring were imported from the custom CQA library, as given in Step 2 (Figure 4A). Each library item provides peptides’ neutral mass and retention time information for accurate peptide detection. In addition, this targeted method is equipped with a relative abundance measurement tool to calculate relative %modification levels of CQAs using single or multiple charge combined ion responses (Figure 4B). Customizable limit settings for %modifications were applied when visualizing the levels of oxidized peptides for easy identification of samples surpassing the standard oxidation levels. The example in Figure 4C demonstrates HC:T21 DTMLISR native and oxidized peptides after limit check using an arbitrary number for the threshold to demonstrate feature capability. Both oxidative stress conditions shown here trigged the limits for modified HC:T21 peptides and were flagged in yellow and red reflecting the preset warning (at 10%) and error limits (at 50%) respectively. The information gathered in each attribute monitoring step can be documented through the automated reporting feature and used in process development of therapeutic proteins.
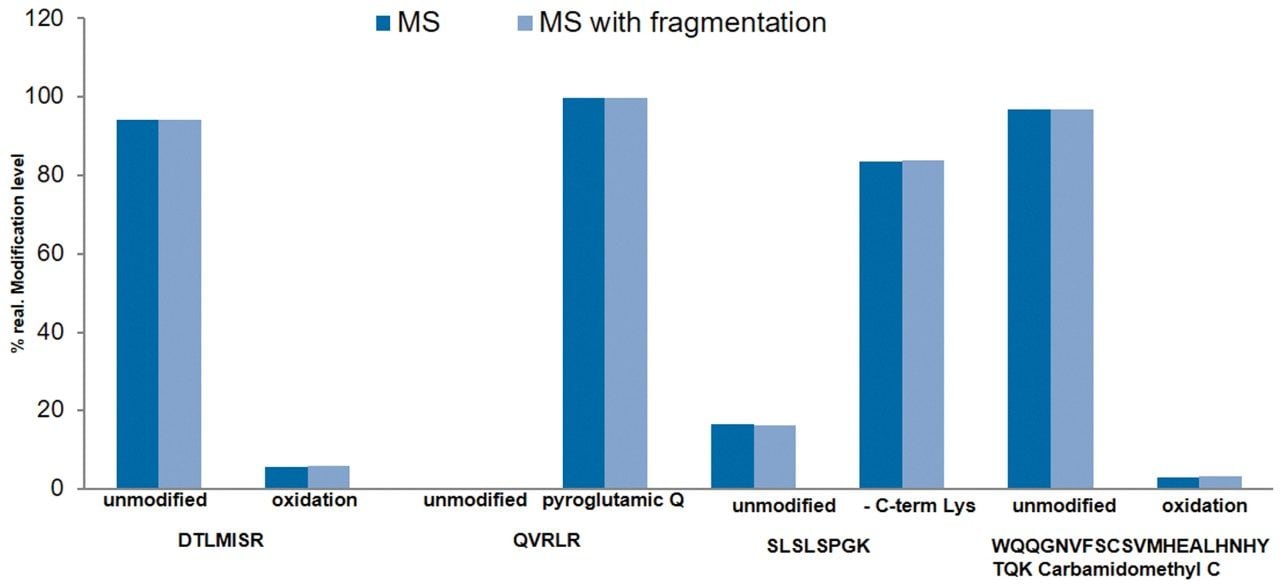
Most peptide monitoring assays use MS only acquisition to screen samples for a list of known target peptides to achieve robust quantification. However, the DIA acquisition with fragmentation spectra can contribute to identification and sequence verification of differentially expressed peaks in mAb samples. The ability to maintain consistent %modification measurements in both MS and MS with fragmentation methods is an important feature that demonstrates data integrity of the platform. In here, the data acquired for mAb control samples using both acquisition methods indicate comparable %relative modifications for the given peptide attributes. Therefore, both methods can be used for routine peptide monitoring (Figure 5) without compromising MS-based %modification levels.
The BioAccord System controlled by compliant-ready UNIFI Software is purposefully designed with integrated methods for routine therapeutic protein analysis. It can perform routine peptide characterization and accurate mass screening for CQA monitoring. The seamless integration of the two peptide analyses through the custom library feature is an effective way of transferring knowledge from characterization to monitoring that can be deployed across different analytical laboratory settings.
Further, the SmartMS System with intuitive informatics control (with promoted tune parameters, automated instrument health status monitoring, and calibration) provides a robust MS system suitable for users with any skill level.5
720006602, June 2019