This application brief demonstrate the performance of the Xevo G2-XS QTof platform for intact level quantification of the trastuzumab monoclonal antibody and the use of the UNIFI Scientific Information System for rapid data processing and analysis.
In this study, a monoclonal antibody is quantified at the intact level through integration of extracted ion chromatograms (XICs).
The discovery and development of biotherapeutic modalities of increasing complexity and high molecular weight creates new quantitative and qualitative analytical challenges. Intact protein quantification using a high resolution mass spectrometer (HRMS) is increasingly performed in discovery bioanalytical labs. Intact mass analysis offers direct quantitative measurement of the whole molecule potentially including sub-species and variant information, as well as providing information about molecular stability and sites of modification. In this technology brief, intact quantification of trastuzumab is demonstrated using the Xevo G2-XS QTof HRMS and the UNIFI Scientific Information System. Quantitative performance attributes including sensitivity, accuracy, and linear dynamic range are presented.
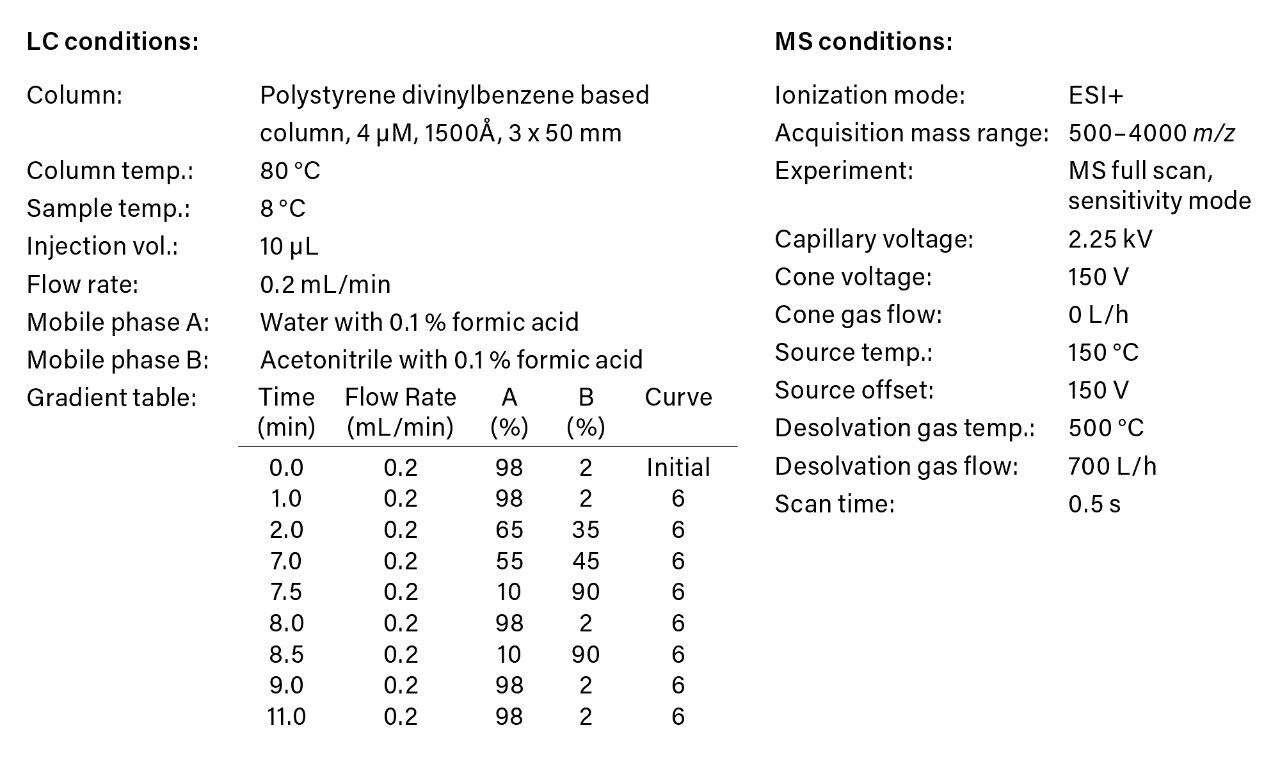
A dilution series of trastuzumab prepared in 0.1% formic acid in the presence of 0.1 mg/mL BSA carrier protein was analyzed using the ACQUITY UPLC H-Class Bio System and Xevo G2-XS QTof. LC-MS acquisition conditions are summarized in Figure 1. The data was processed post acquisition in UNIFI, a next generation informatics platform offering a friendly user interface for data processing, review, and reporting.1 Here, we show a complete quantitative workflow for processing of intact protein data. By scrolling through the workflow, one can quickly review the analysis of a batch of samples including various quantitative performance attributes (Figure 2).
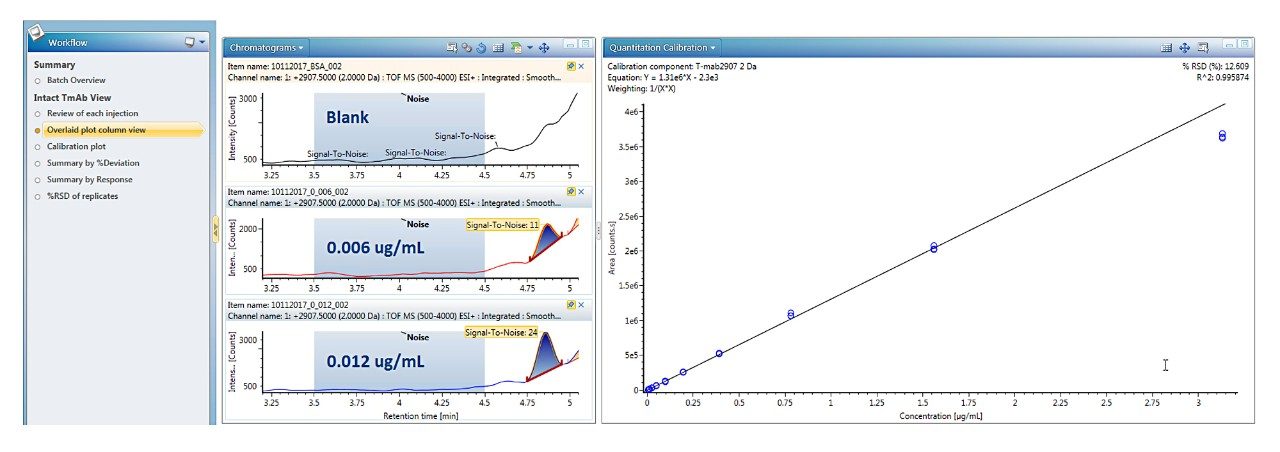
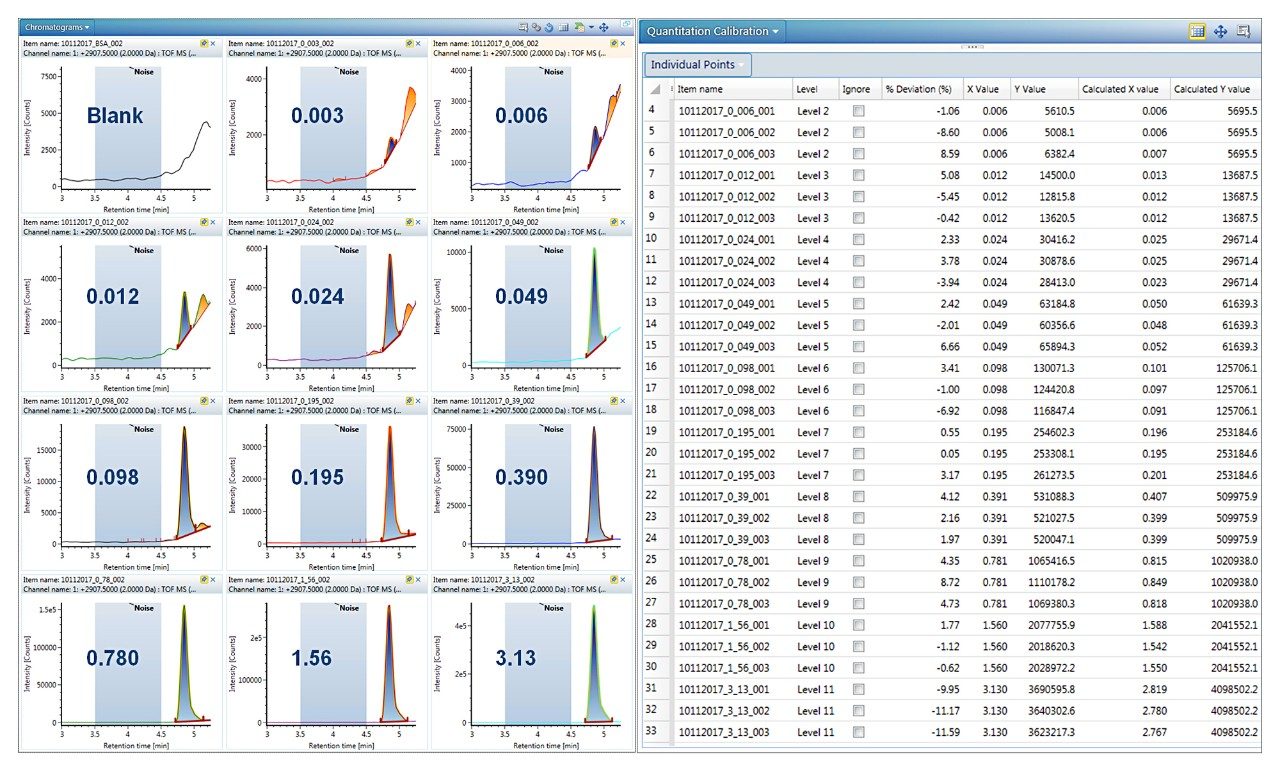
Data processing for quantification of intact proteins was performed by automatically generating and summing extracted ion chromatograms (XICs) from the dataset. Single or multiple peaks can be selected from one or multiple charge states using a mass tolerance window. A detailed investigation of these options on quantitative performance including LLOQ is summarized in a recent Waters Application Note (720006222EN).2 For the present data set, a mass tolerance window of 2 Da for both a single peak at m/z 2907 and the sum of 9 GF0/GF1 peaks across multiple charge states were found to produce the same quantitative results; single peak data is reported here.
Trastuzumab eluted at 4.9 min, as a symmetrical peak, with a linear dose response curve for concentrations ranging from 0.006 to 3.13 μg/mL or 2.7 orders of magnitude. Using 1/X2 weighting, the linear regression produced an R2 = 0.996 (Figure 2). The LLOQ was 0.006 μg/mL from triplicate injections all with a signal-to-noise ratio >10. The calibration table in Figure 3 shows that calculated %accuracy for triplicate injections across all concentrations is within 15%, with no data outliers (Figure 3).
HRMS with its extended mass range makes it an ideal platform for intact quantification of larger proteins such as monoclonal antibodies. In the present study, a monoclonal antibody, trastuzumab, was quantified at the intact level through integration of XICs, which follows common practice in small molecule and peptide bioanalysis. The Xevo G2-XS QTof (HRMS) platform was shown to provide sensitive detection, achieving a LLOQ of 0.006 μg/mL or 60 pg load on column, and a linear dynamic range of 2.7 orders of magnitude that is suitable for routine intact monoclonal antibody quantifications.
UNIFI offers direct data import of MassLynx .raw data files. Instructions for importing data are provided as follows:
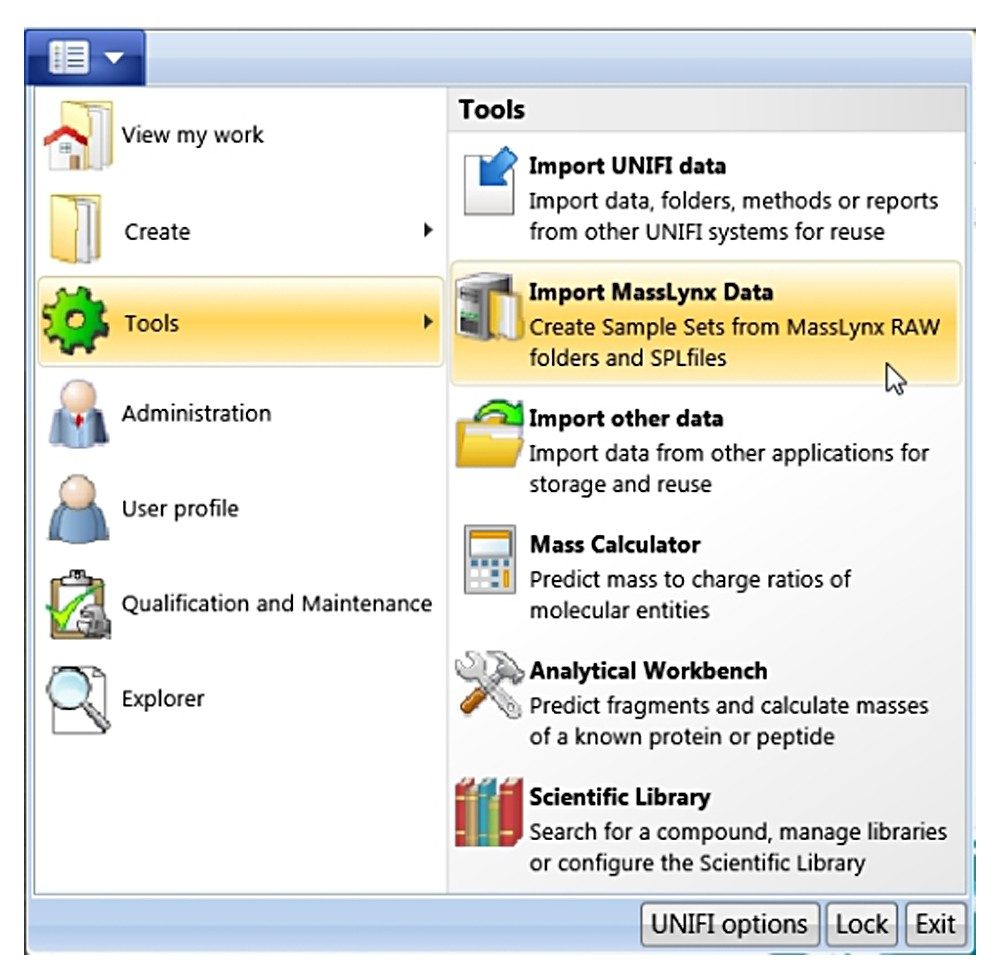
Click the drop-down arrow of the main menu, a panel shown in Figure A will appear. Choose “Tools”, “Import MassLynx data”. A new dialog “Create Import Task” shown in Figure B will appear. From this dialogue, one can choose either “MassLynx raw folders” for importing data only (*.RAW), or “MassLynx SPL files” for importing through a MassLynx sample list file (*.SPL). The advantage of using the sample list file type is that the injection sequence of the imported samples is the same as the original.
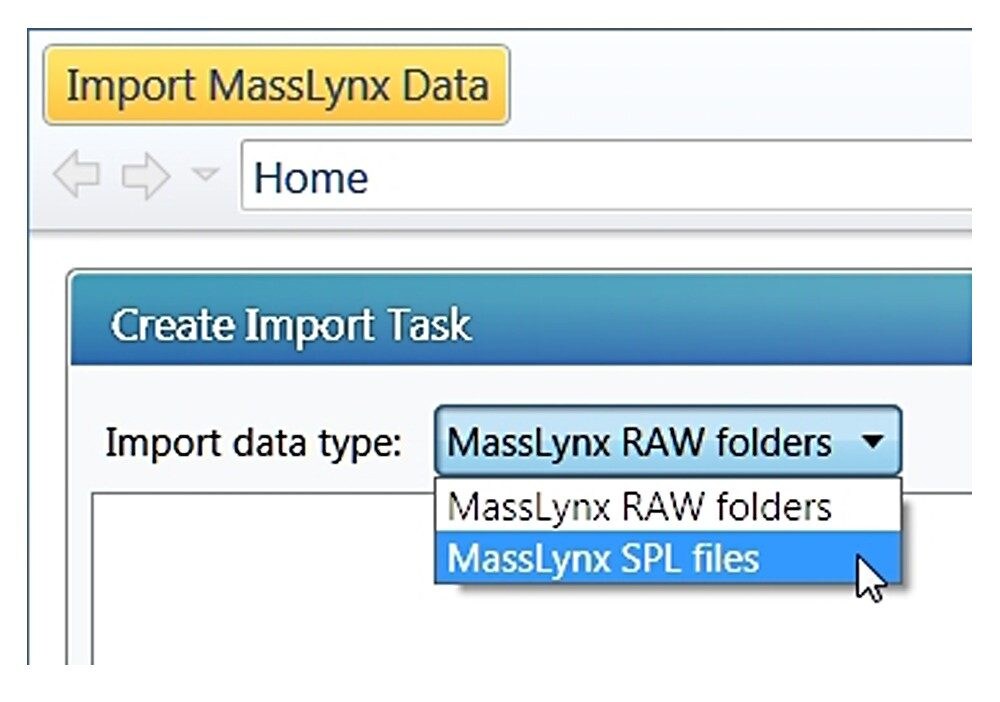
With the “MassLynx SPL file” import type chosen, one can import directly from a MassLynx.PRO project folder or from copied data. Note: In the case of copied data, the sample list file .SPL needs to reside in a folder called “SampleDB”, and .raw files need to reside in a folder called “Data” as shown in Figure B (right). In this way, the sample list will correctly associate each entry with a data file.
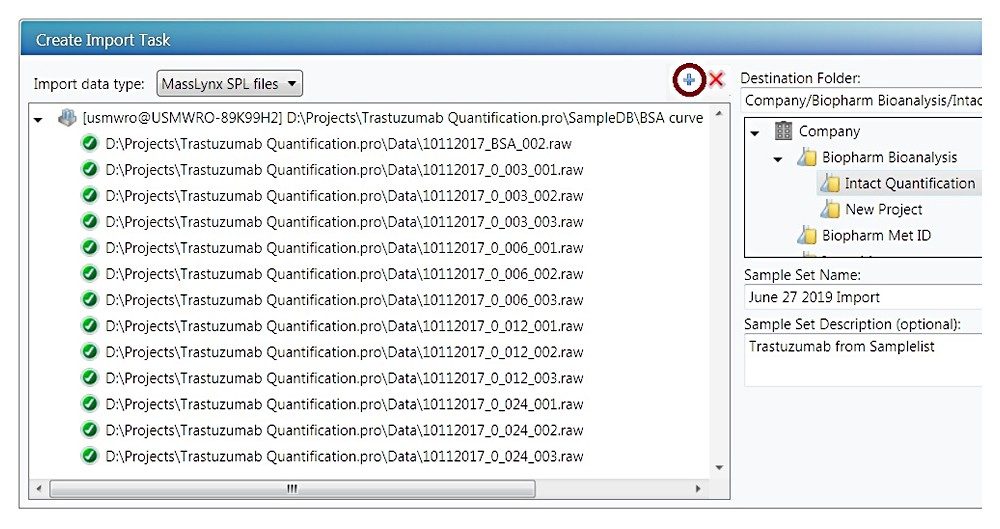
With choosing the “MassLynx SPL data files”, click the “+” button, a file browsing window will appear (not shown). Browse to the .SPL file location and click OK, the sample list file and corresponding .raw data files will populate the left panel of Figure C. Choose a destination folder and enter the sample set name, pressing the “Create UNIFI Sample Set” button will start the import. A green bar shown in the right panel shows the progression of the import.
After the import is completed, the files will appear in UNIFI Explorer with the sample set format. The user can then create an analysis and perform data processing.
720006379, September 2018