For research use only. Not for use in diagnostic procedures.
This is an Application Brief and does not contain a detailed Experimental section.
This application brief evaluates the effect of SONAR quadrupole window size on sensitivity when narrowing the window to gain higher selectivity in discovery omics analysis.
SONAR DIA is for highly selective fragmentation spectra resulting in more efficient database searching.
The accurate identification of peptides, proteins, lipids, and small molecule metabolites in omics studies requires the generation of high quality precursor MS and product ion MS/MS data. Accurate mass data independent acquisition (DIA) LC-MS is frequently employed in metabolite identification and omics studies, providing a broad coverage of MS and MS² information simultaneously during an analytical batch.1–2
DIA strategies implemented on QTof platforms have been demonstrated previously. For example, a highly specific DIA approach with use of a rapidly scanning quadrupole (SONAR) has shown to provide high specificity that is compatible with fast chromatographic separation.³ Quadrupole separation prior to fragmentation provides specificity allowing fragment ions to be readily correlated with their associated precursors, highlighting the advantages over techniques which do not use a quadrupole or alternative means of MS1 isolation. SONAR is shown to provide increased MS² spectral quality and ultimately improved database search results.
Narrowing the quadrupole spectral window, provides cleaner spectra and therefore more efficient compound library searches (720006036EN). A narrow quadrupole window requires a fast scanning quadrupole to provide a sufficient number of points across an LC peak, to retain quantitative accuracy; however, this increased duty cycle can give rise to a reduction in sensitivity.
SONAR is a novel DIA acquisition method that employs a narrow pass filter and rapid scanning quadrupole. The SONAR acquisition window can be varied from 5–35 Da allowing for a smaller or larger selection of ions prior to entering the collision cell. The effect of the quadrupole window on analytical system sensitivity was evaluated by analyzing the lipid profile of human serum using a reversed phase UPLC-MS method over 15 minutes with the ACQUITY UPLC I-Class System and Xevo G2-XS QTof.
Data were collected in positive ion mode using a SONAR quadrupole window which varied between 10 and 35 Da as well as an unbiased DIA acquisition (MSE) and full scan MS over the TOF mass range of 100–1000 Da. The resulting data were processed and database searched using Progenesis QI Software. Results from the analysis allowed for the low energy peak count of each sample, average low energy peak count and total ion current intensity to be determined.
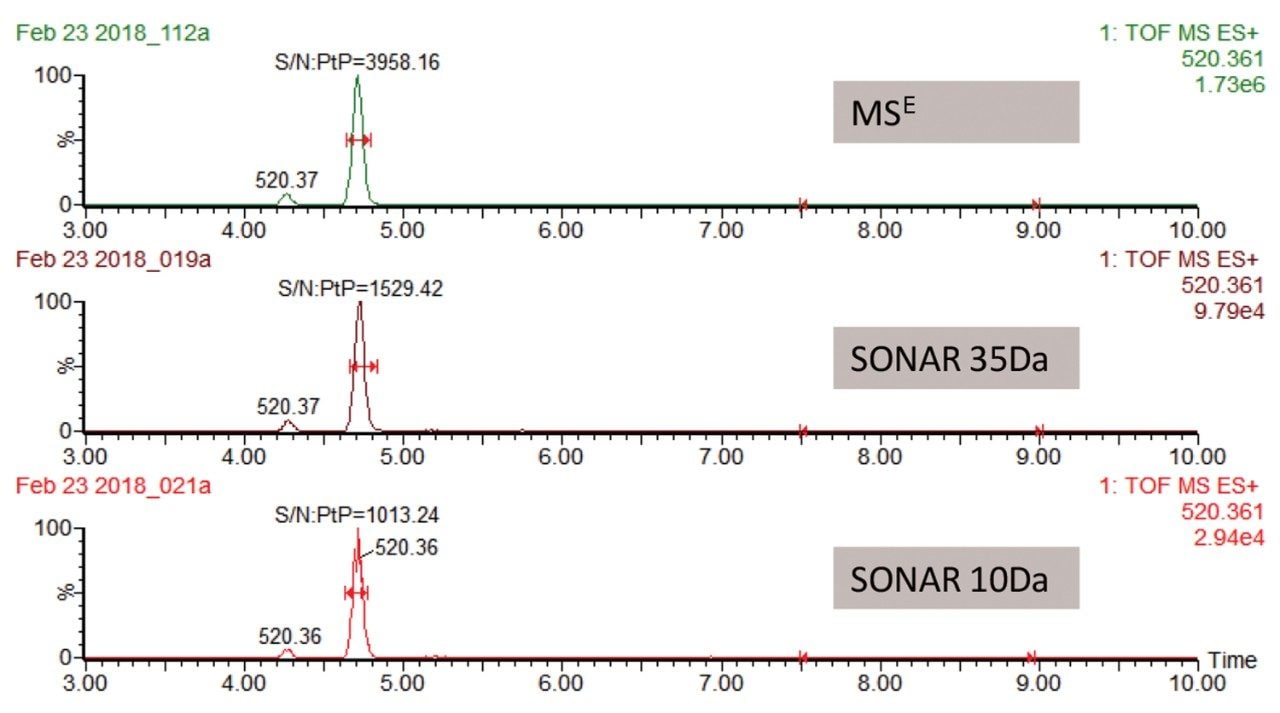
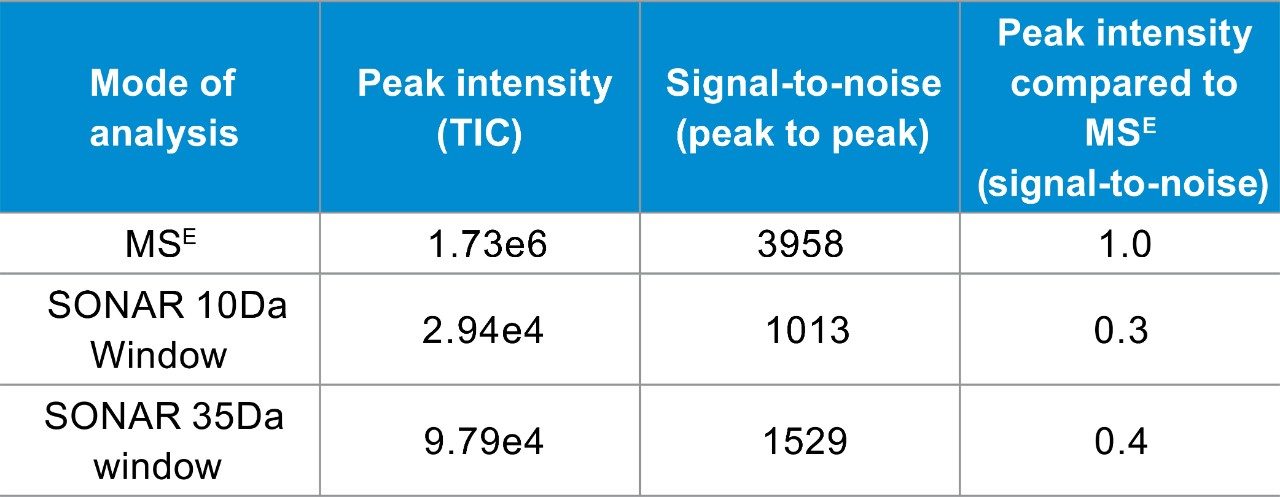
The data displayed in Figure 1 illustrates changes in the lipid assay sensitivity for SONAR based quadrupole window sizes of 10 and 35 Da in addition to MSE. From this data, it is clear that the MSE mode of analysis provided the highest response (based on overall total ion count (TIC)) with a maximum intensity of 1.73e6. The SONAR analysis using a 10 Da window produced a peak response of 2.94e4, while the 35 Da window produced a peak response of 9.79e4 (Table 1). The signal-to-noise for each peak was then compared to evaluate the usable sensitivity of each acquisition mode. The data showed that while full scan MS and MSE acquisition modes were virtually equivalent, SONAR was shown to provide a signal response that was only 2.4 times lower than full scan MS mode for equivalent sample loading. Observing only a small reduction in signal response can be attributed to the substantial noise reduction observed with SONAR data, due to the enhanced specificity provided by the scanning quadrupole.
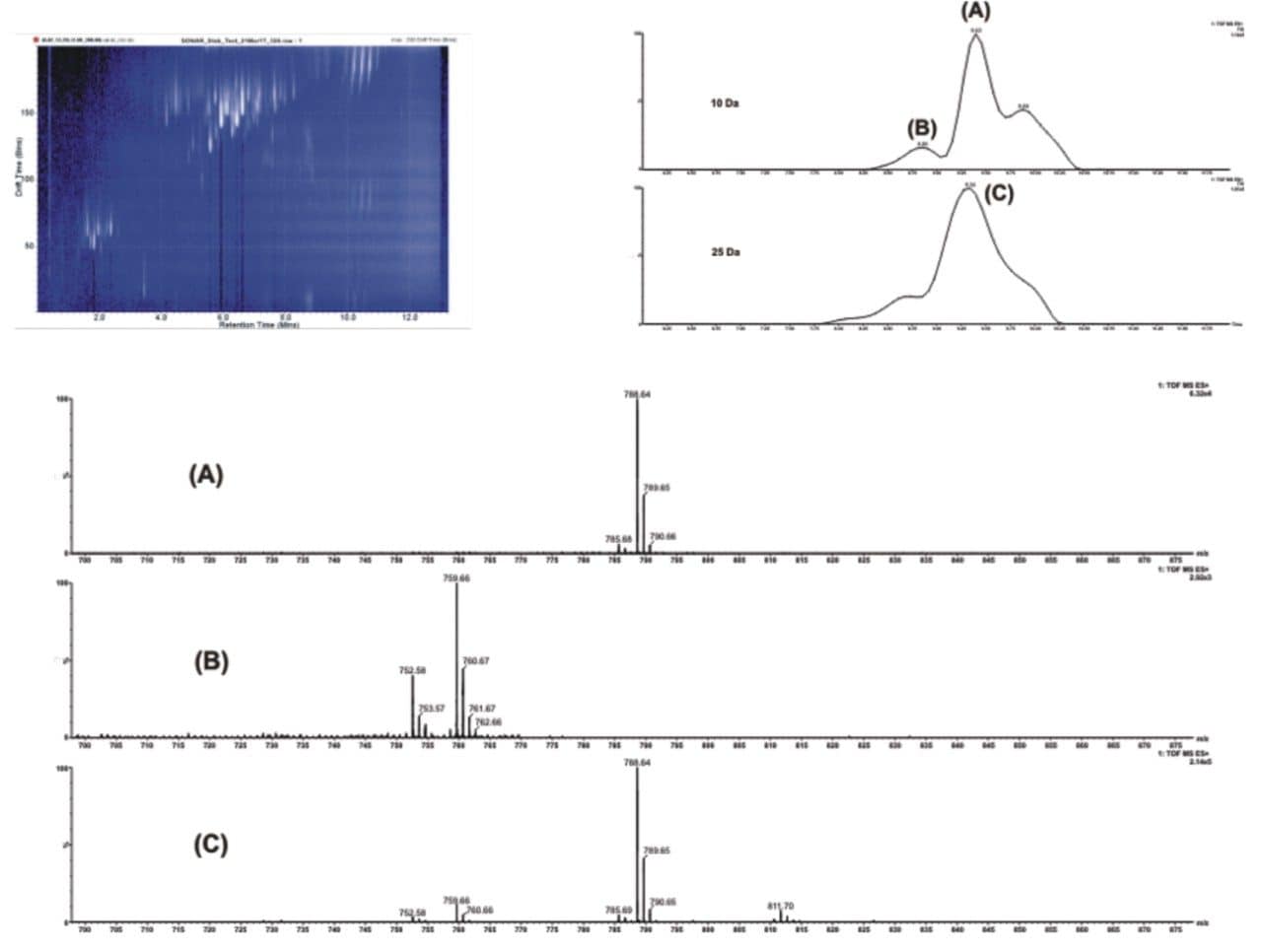
The benefits of the 10 Da window size is illustrated in Figure 2 which shows increased selectivity when using narrower quadrupole settings. Comparing a region of interest highlights the gains in selectivity. Extracted spectra show cleaner spectra with discrete peaks due to enhanced specificity, with lipids of interest clearly distinguished.
SONAR is a novel DIA acquisition for a hybrid quadrupole time-of-flight mass spectrometer (QTof), which employs a narrow pass, fast scanning quadrupole filter. This mode of operation produces cleaner MS² spectra resulting in more efficient database searching. When using a 35 Da window the SONAR mode of analysis showed a 2.4 fold reduction in detection sensitivity compared to full scan MS based on signal-to-noise (peak to peak). The impact of the quadrupole filter size on MS sensitivity and peak response has been evaluated from 10–35 Da using a lipid profiling method. The results showed that the 10 Da window delivered clear, superior LC-MS data quality while maintaining sufficient sensitivity for quantitative measurements.
720006331, July 2018