This is an Application Brief and does not contain a detailed Experimental section.
This application brief utilizes Skyline SRM/MRM Library Explorer and Waters Xevo TQ-S formats to enable intuitive, prompt, and seamless development of multiple SRM/MRM peptide transitions and methods from high-density proteomics LC-MS experiments.
Utilize data from cross platform DDA and DIA proteomics discovery experiments to quickly and easily design robust, highly sensitive, and highthroughput validation MRM assays with Xevo TQ-S.
Targeted LC-MS/MS SRM/MRM validation has the potential to bridge protein biomarker discovery and validation. Candidate proteins and peptides can be retrieved from one or multiple identification repositories and libraries. Typical bottlenecks observed are the selection of the appropriate proteotypic peptides for SRM/MRM quantitation and the development of the associated LC-MS methods for triple quadrupole mass spectrometers. Software plays a vital role in selecting the correct peptides, especially in the case of complex experiments, to measure as a substitute for the target protein. This transitions into measurement of target peptides, where specific optimizations for each are typically performed. Skyline Discovery Library Explorer is an open source document editor for creating and analyzing targeted proteomics experiments.1 In this technology brief we describe the use of Skyline for the peptide and transition selection process from libraries for method creation for a Xevo TQ-S Mass Spectrometer.
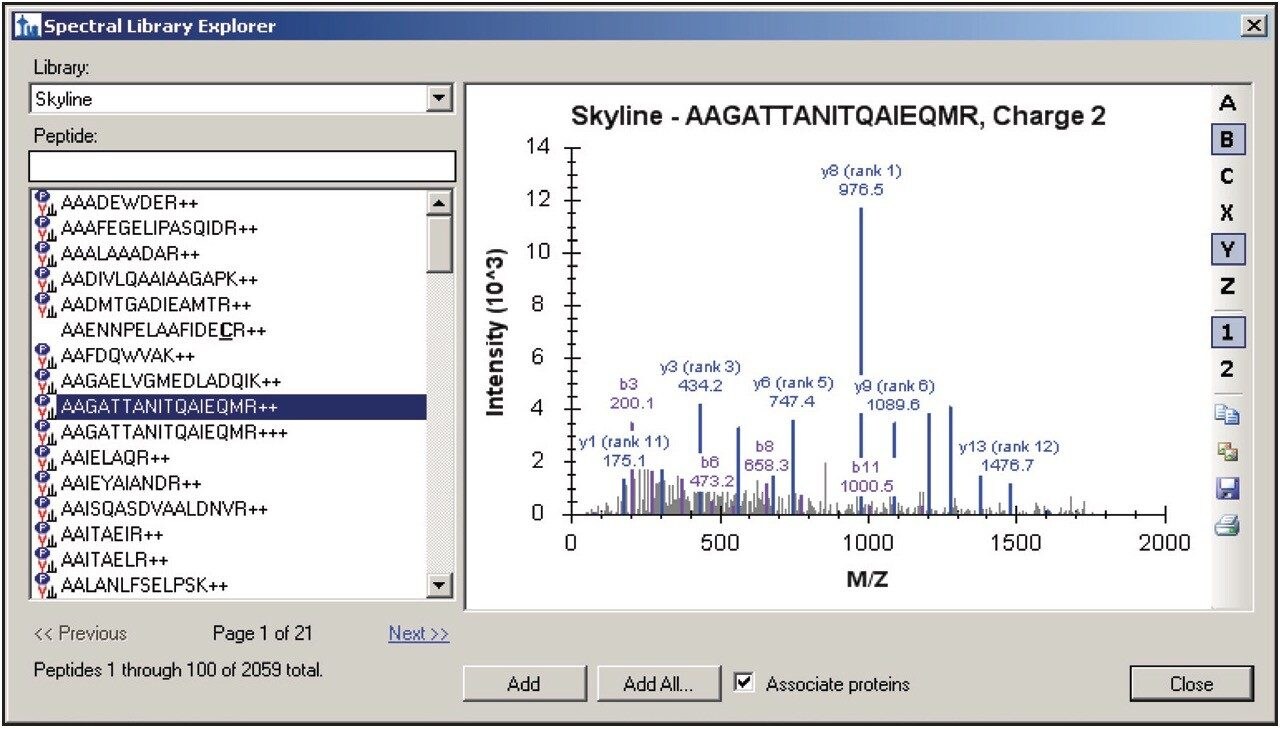
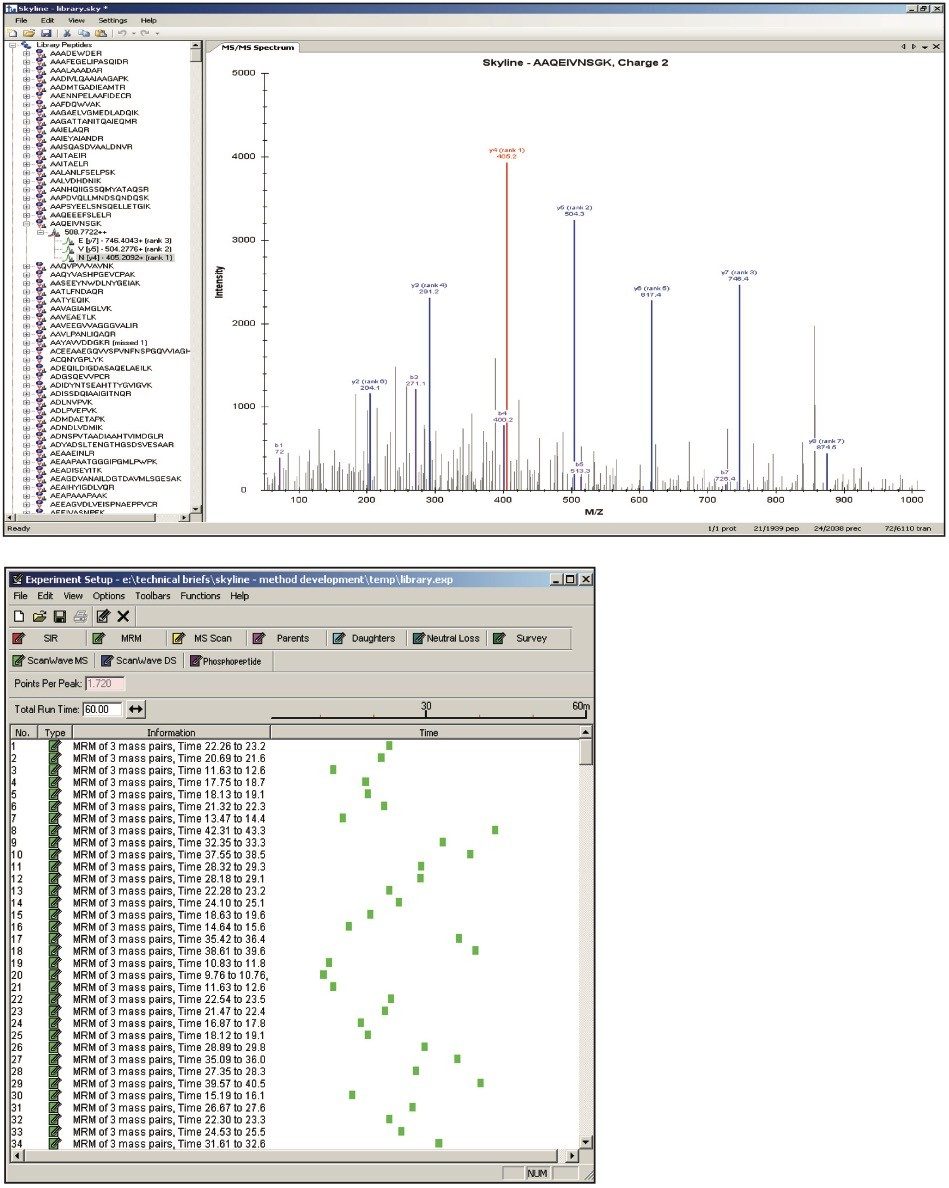
A peptide library was created from the search results of a DDA LC-MS/MS discovery experiment and a common protein database search engine that analyzed tryptically digested cytosolic E.coli spiked with four protein digest standards. One or more peptides, shown in Figure 1, were selected and automatically associated with their parent proteins for SRM/MRM method creation. Next, Skyline determined and presented the best candidate transitions. The three most abundant product ions from which a Xevo TQ-S method can be automatically built are shown in the upper and bottom panes of Figure 2. In this instance, a scheduled SRM/MRM method was created using the built-in retention time model of Skyline and empirical observations as input/landmarks for retention time prediction.
The results are summarized in Figure 3, which shows the summed SRM/MRM chromatograms of two transitions per peptide. Also shown are the responses of two background, confirmatory E. coli peptides and those of two of the target components. In this example, peptides from yeast alcohol dehydrogenase (ADH) and enolase, were spiked and detected with confident signal-to-noise levels at the 1.25 attomole level on-column.
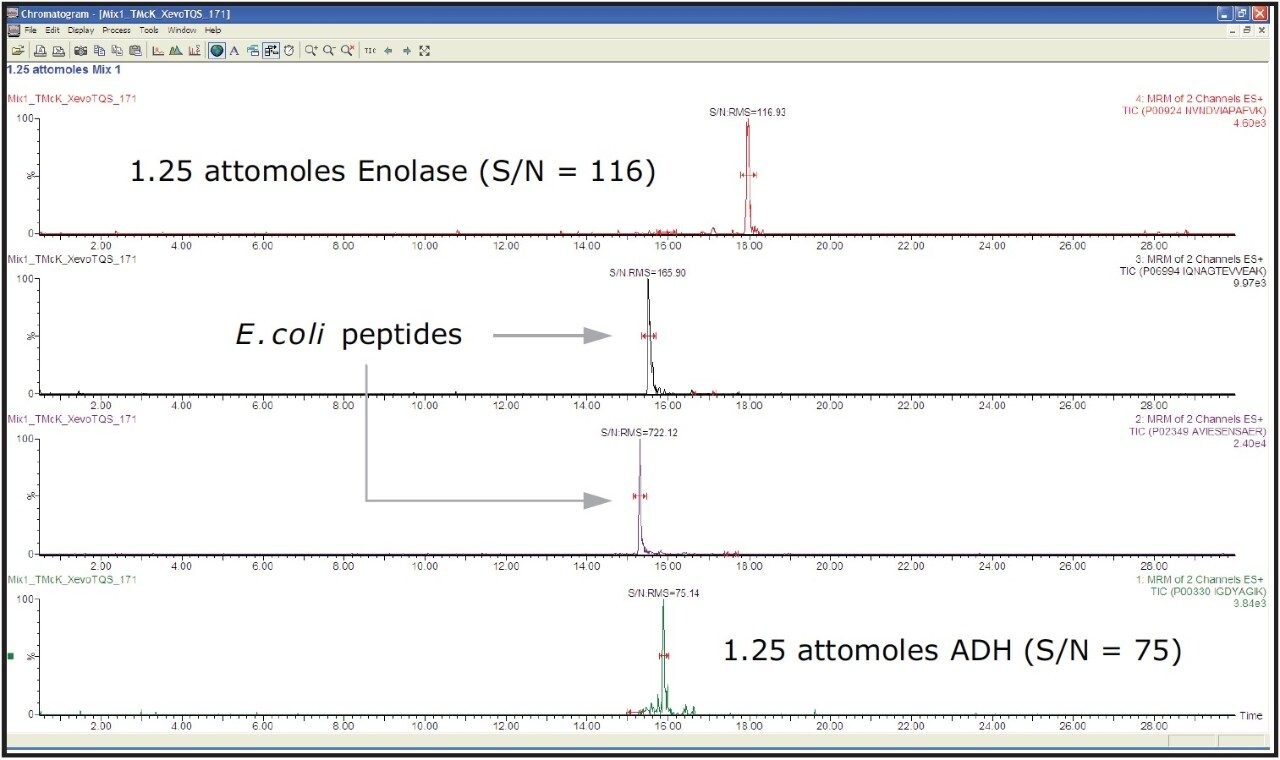
The SRM/MRM method development of target standard peptides and background E.coli peptides for Xevo TQ-S has been demonstrated using Skyline and discovery search results, libraries, and repositories.
720004345, May 2012