This work evaluates the use of targeted LC-MS analysis using multiple reaction monitoring (MRM) for fast and sensitive monitoring and quantification of degradations in mAb digests. This approach, using UPLC with a sensitive tandem quadrupole MS, can also be adapted for monitoring and quantification of other PTMs and degradations such as N-linked glycosylation. This solution offers a significant improvement in speed, specificity, and sensitivity of mAb analysis and has the potential to expedite the research and development of protein drugs.
Post-translational modifications (PTMs) and chemical degradations impact the efficacy and safety of therapeutic proteins and peptides. Therefore, they have to be effectively controlled and monitored during production, formulation, and storage. Liquid chromatography (LC)-based peptide mapping with ultra violet (UV) and/or mass spectrometry (MS) detection is a method of choice for characterization of PTMs and degradations.1
However, higher throughput and more sensitive methods are needed for routine monitoring and quantification of protein drugs during development and manufacturing. For example, in protein formulation development hundreds of samples are typically analyzed to insure purity and stability. Although intact protein mass measurement (including limited proteolysis followed by LC-MS analysis) can be used for fast evaluation of large mass-shifted PTMs such as glycosylation, it does not provide information on modification sites and has limitations for detection of modifications with small mass shifts,2, 3 such as asparagine (N)-deamidation, aspartic acid (D)-isomerization, and methionine (M)-oxidation.
Monoclonal antibodies (mAbs) are an important class of biotherapeutics. They are susceptible to a number of PTMs and degradations common to other classes of therapeutic proteins. Peptide mapping studies3, 4 demonstrate that PTMs and degradations, such as glycosylation, deamidation, and oxidation, only occur in specific motifs of a mAb. This application note evaluates the use of targeted UPLC-MS analysis using multiple reaction monitoring (MRM) for fast and sensitive monitoring and quantification of degradations in mAb digests.
As a proof of concept, we performed a UPLC/MRM study with the goal of confirmation and quantification of site-specific M-oxidations in a recombinant IgG1 tryptic digest previously identified by UPLC/MSE peptide mapping studies.3, 4
Both oxidized and unmodified M-containing tryptic peptides were unambiguously determined by multiple MRM transitions, and relative quantification data were obtained for a low-abundance M-oxidized peptide.
The large dynamic range and linearity of the combined ACCQUITY UPLC/Xevo TQ MS System5,6 used for this UPLC/MRM assay offers a significant improvement in speed, specificity, and sensitivity of mAb analysis and has the potential to expedite the research and development of protein drugs by saving both time and analytical resources.
|
LC system: |
ACQUITY UPLC |
|
Column: |
ACQUITY UPLC BEH 300 C18 a1.7 μm, 2.1 x 150 mm (p/n 186003687) |
|
Column temp.: |
60 °C |
|
Flow rate: |
300 μL/min |
|
Injection vol.: |
10 μL (partial loop injection mode) |
|
Mobile phase A: |
0.1% FA in water |
|
Mobile phase B: |
0.1% FA in ACN |
|
Gradient: |
0-40% B in 60 min |
|
MS system: |
Xevo TQ MS |
|
Ionization mode: |
ESI+ |
|
Acquisition mode: |
MRM |
|
Capillary voltage: |
3.5 kV |
|
Cone voltage: |
35 V |
|
Mass window: |
1 Da (unit mass resolution) |
|
Dwell time: |
5 ms |
Synthetic peptides DIQMTQSPSSLSASVGDR (purity 80%) and DIQMoxTQSPSSLSASVGDR (purity 87%, Mox-oxidized methionine) were purchased from Biomatik Corporation (Canada). The IgG1 antibody trastuzumab digest was prepared by a RapiGest-assisted 4-h trypsin digestion protocol.7,8
In previous LC-MSE peptide mapping studies3,4 we detected two primary M-oxidation sites from the antibody. One site is located in heavy chain tryptic peptide HT21 (DTLMISR, ~4% oxidation) and the other in light chain tryptic peptide LT1 (DIQMTQSPSSLSASVGDR, <0.5% oxidation). Because of the complexity of the mAb digest, low-abundance oxidized peptide peaks are often obscured by more dominant peptides in both LC/UV and LC-MS analysis.
Here, we demonstrate that LC/MRM analysis provides enhanced selectivity and can detect M-oxidation of peptides at low-abundance. The method is potentially applicable for monitoring other PTM in protein drugs as well.
Figure 1 contains the MRM chromatograms of unmodified and oxidized LT1 eluted from the antibody tryptic digest. As expected, oxidized LT1 (at 19.35 min) eluted earlier than unmodified LT1 (at 23.29 min) due to reduced hydrophobicity upon M-oxidation. The use of three MRM transitions (see Figure 1 for details) clearly confirms the identity of the peptides, demonstrating that the use of multiple MRM transitions in this manner has advantages over single ion monitoring (SIM) or extracted ion chromatogram (XIC) techniques for monitoring protein PTMs from complex samples such as protein tryptic digests. Very limited in-source M-oxidation9 (<0.01%) was observed in this experiment (signal present at retention time identical to non-oxidized peptide peak).
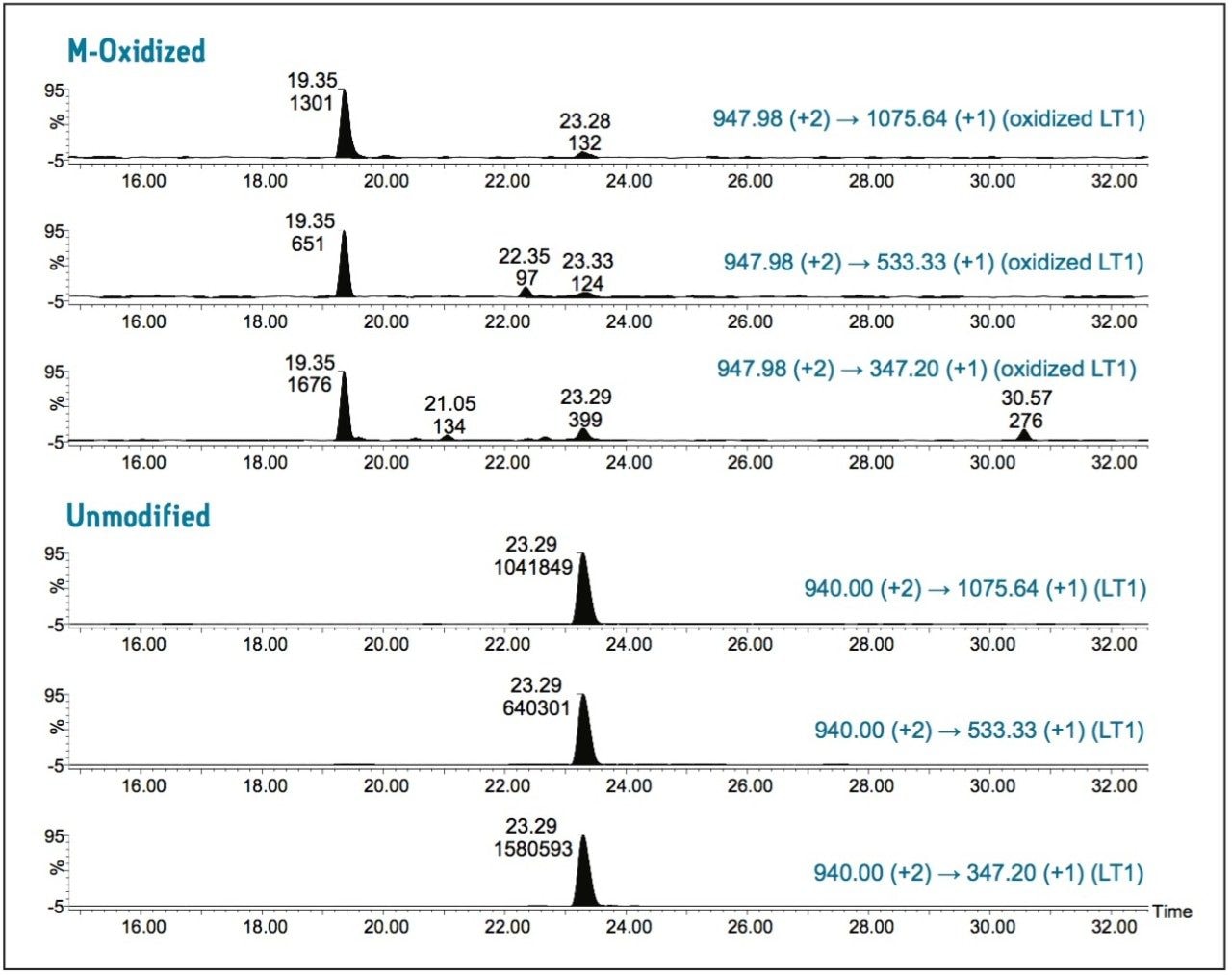
The same relative concentration (~0.1%) of oxidized LT1 was obtained when quantifying by integrated peak areas of either corresponding transition pair. The relative standard deviation (RSD in %) for integrated peak area of unmodified LT1 is 2.7%; the RSD for low-abundance oxidized LT1 is 23.9% (calculated from 3 repeated injections). The relatively large RSD for low-abundance oxidized LT1 is due to the MS response variation of the minor signal. The MRM transition of precursor to fragment y3 (374.20 Da) has the highest intensity for both oxidized and unmodified LT1. Quantification using peak areas of this transition provides the highest sensitivity and accuracy.
Next, we tested the dynamic range and quantification linearity of the method using synthetic oxidized and unmodified LT1. The transitions 947.98 Da to 347.2 Da (for oxidized LT1) and 940.0 Da to 347.2 Da (for unmodified LT1) were selected for this study because of their high MS response.
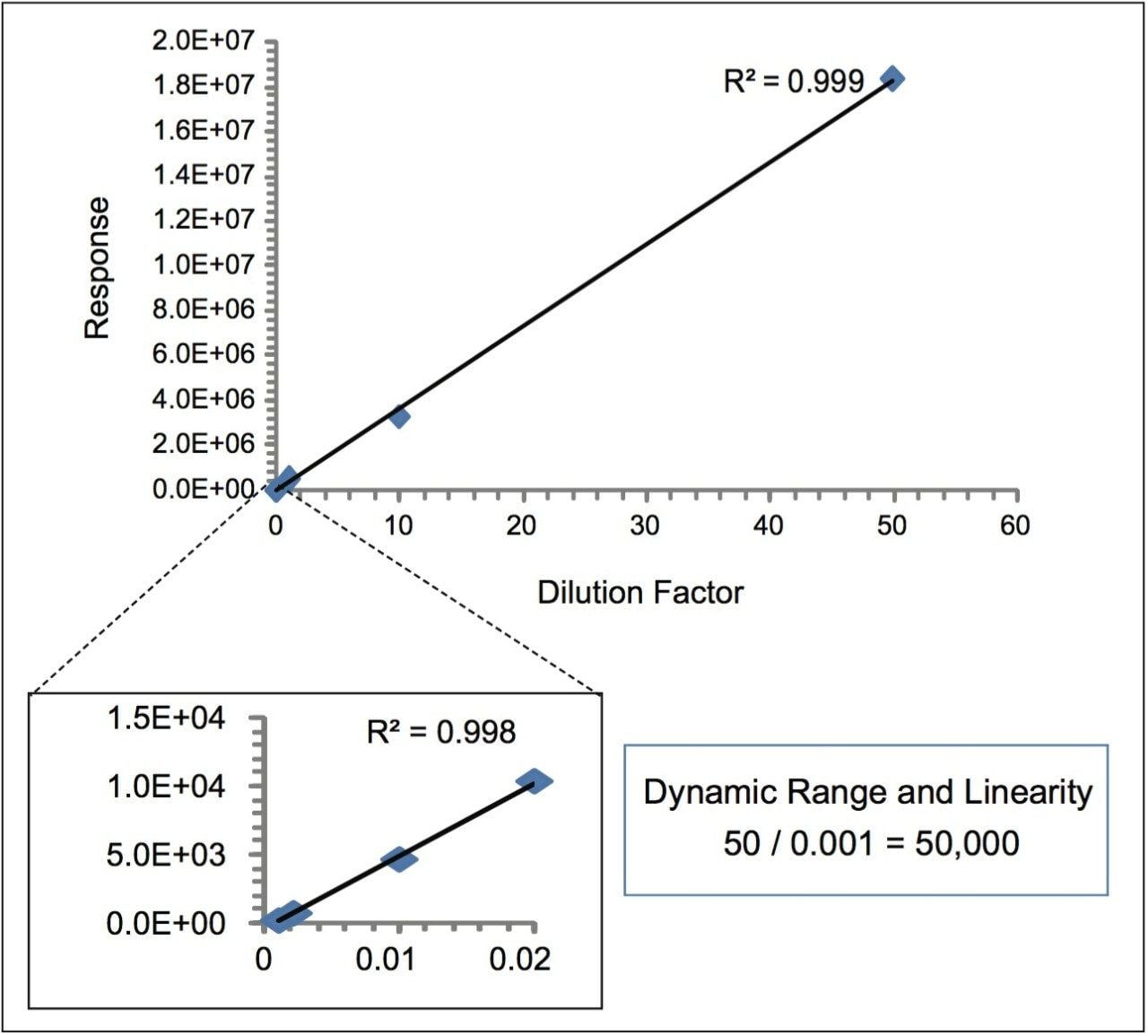
Four orders of dynamic range and linear response (from low fmole to nmole) were observed for both unmodified and oxidized LT1. This indicates that M-oxidation with relative concentration as low as 0.01% could be detected and quantified by this method. The MRM response of LT1 at different dilutions and concentrations of the stock (with dilution factors of 0.001, 0.002, 0.01, 0.02, 1, 10 and 50) was plotted in Figure 2. The estimated limit of detection (LOD) and limit of quantification (LOQ) were below 50 fmole.
A UPLC/MRM assay was developed for quantification of site-specific M-oxidization in a recombinant monoclonal antibody. The method offers the following benefits:
The technology demonstrated here can also be adapted for monitoring and quantification of other PTMs and degradations such as N-linked glycosylation. Unlike traditional MRM methods, no internal standards are required for relative quantification of PTMs and degradations in therapeutic proteins.
The technology could be targeted for high throughput monitoring of critical attributes in biotherapeutics from batch to batch.
720003780, October 2010