
For research use only. Not for use in diagnostic procedures.
This application note investigstes rapid profiling of lipids originating from human plasma derived from breast cancer and control samples.
The metabolic and lipidomic profiling of biofluids and tissues has been proven to provide new insights into disease mechanism and progression.1 Early pilot studies with small sample batches not only demonstrated the capability of the “omics” approach but also revealed the issues arising from small cohort studies, in terms of misleading results due to a lack of population, gender, and age coverage.2 Large scale metabolic phenotyping studies require the ability to perform accurate and reproducible analysis of large sample cohorts and can often require several batches of analysis to complete.
The analyses of larger cohorts, for example 5000 samples and upwards, require either a large instrument facility or faster methodology.3 Typical LC-MS profiling methods have sample cycle times of 10–20 minutes, thus a continuous analysis based on conventional UPLC for a study cohort of 1000 samples would require several days of instrumentation time. The benefits of rapid LC analysis combined with microbore LC columns for large batch analysis was demonstrated by Wilson et al. for the analysis of a pre-clinical safety assessment study.4 The number of overall detected features with RAMMP can be compromised when compared with UPLC. However, combining the RAMMP methodology with data independent acquisition (DIA) strategies involving an ion mobility (IM) schema results in both high peak capacity and ultimately larger numbers of detected features being reported.
A pooled sample was prepared by combining 10 μL of all samples (breast cancer patients and healthy controls) which were subsequently stored at -20°C until use. Prior to analysis, each sample was subject to precipitation using the ratio of 1:4 with isopropanol (IPA). The sample was then vortex mixed and centrifuged at 13,000 rpm in a microfuge for five minutes. The resulting supernatant layer was removed and transferred to an autosampler vial for analysis.
|
LC system |
ACQUITY UPLC I-Class |
|
Column |
ACQUITY BEH C8 1.0 × 50 mm |
|
Column temp. |
55 °C |
|
Sample temp. |
10 °C |
|
Injection volume |
0.2 μL |
|
Flow rate |
250 μL/min |
|
Mobile phase A: |
(50:25:25) H20:IPA:MeCN w/5 mM ammonium acetate and 0.05% acetic acid |
|
Mobile phase B: |
(50:50) IPA:MeCN w/5 mM ammonium acetate and 0.05% acetic acid |
|
Time (min) |
Solvent composition (%A) |
Solvent composition (%B) |
|
0.05 |
90 |
30 |
|
2.8 |
30 |
90 |
|
3.0 |
0.1 |
99.9 |
|
3.15 |
0.1 |
99.9 |
|
3.7 |
90 |
30 |
|
MS system: |
Synapt G2-Si |
|
Ionization mode: |
ESI+ |
|
Acquisition range: |
50–1200 m/z |
|
Capillary voltage: |
3.0 kV |
|
Collision energy: |
Low energy 5eV, high energy 25eV |
|
Cone voltage: |
30 V |
|
MS software |
MassLynx |
|
Informatics |
Progenesis QI |
The lipidomic analysis of samples arising from large cohorts requires increased analytical throughput. Simply reducing the LC analysis time however, would result in a loss of chromatographic peak capacity. To address these concerns, the separation was geometrically scaled from a 2.1 × 100 mm ACQUITY BEH C8 Column to a narrow bore 1.0 × 50 mm ACQUITY BEH C8 Column, with a mobile phase flow rate of 250 μL/min and an injection volume of 0.2 μL. This approach ensures that the number of column volumes defining the gradient was kept approximately constant between the two separations, resulting in the chromatographic analysis time being reduced from 15 to 3 minutes.The chromatographic performance of the RAMMPlipid methodology was evaluated using the AvantiLipid differential ion mobility mix. The resultingdata displayed in Figure 1, shows that the systemdelivered a high quality separation that wascapable of separating standards in the testmix within the 3.7 minute analysis time.
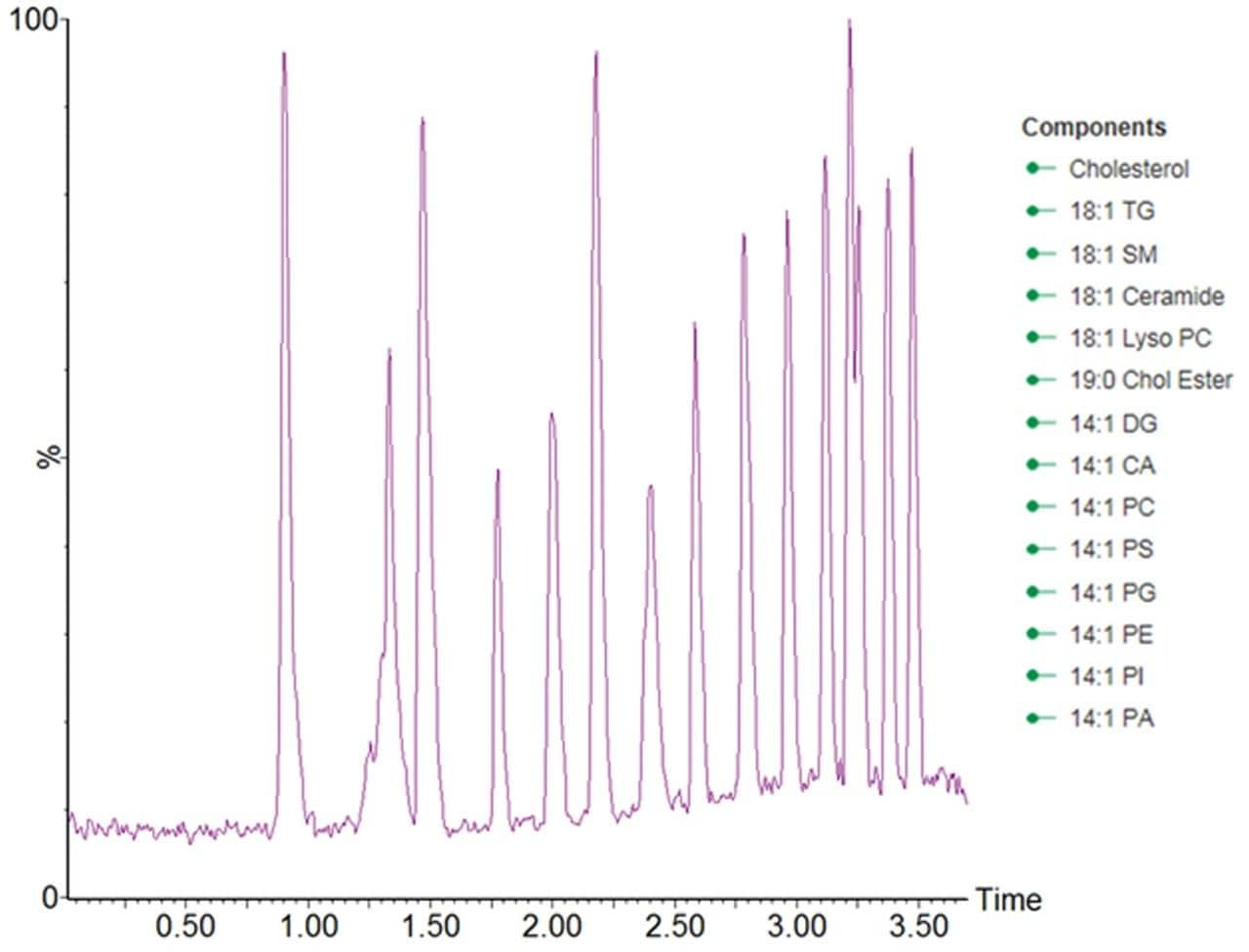
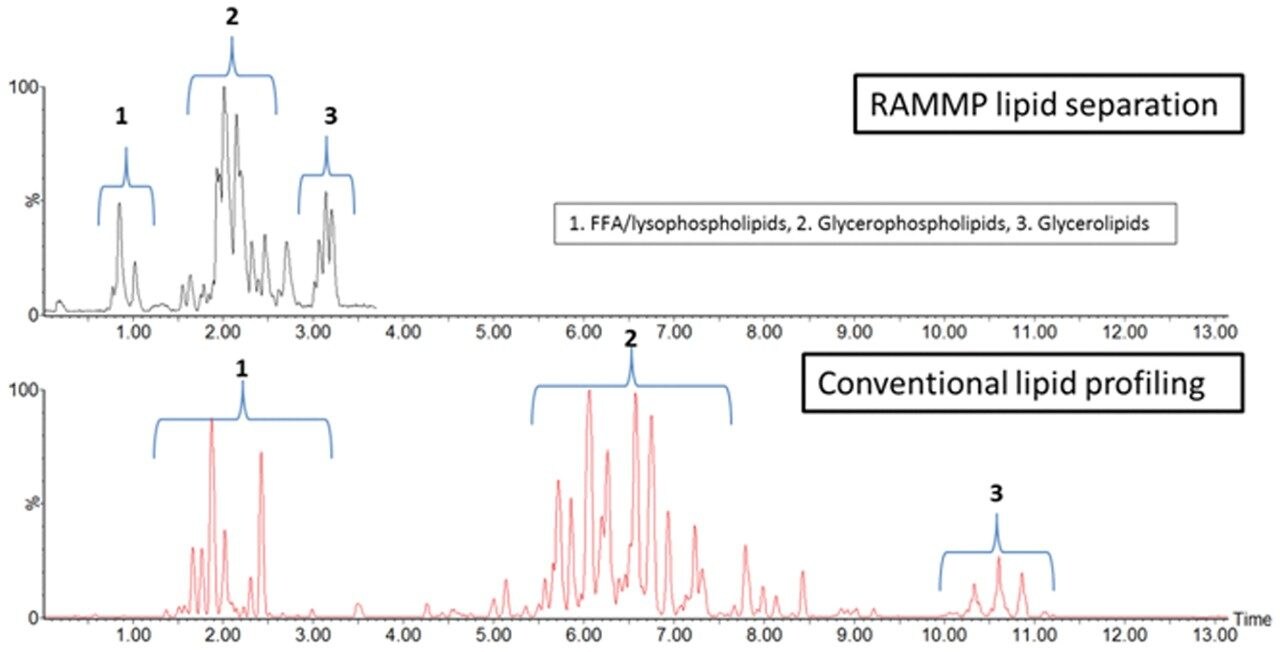
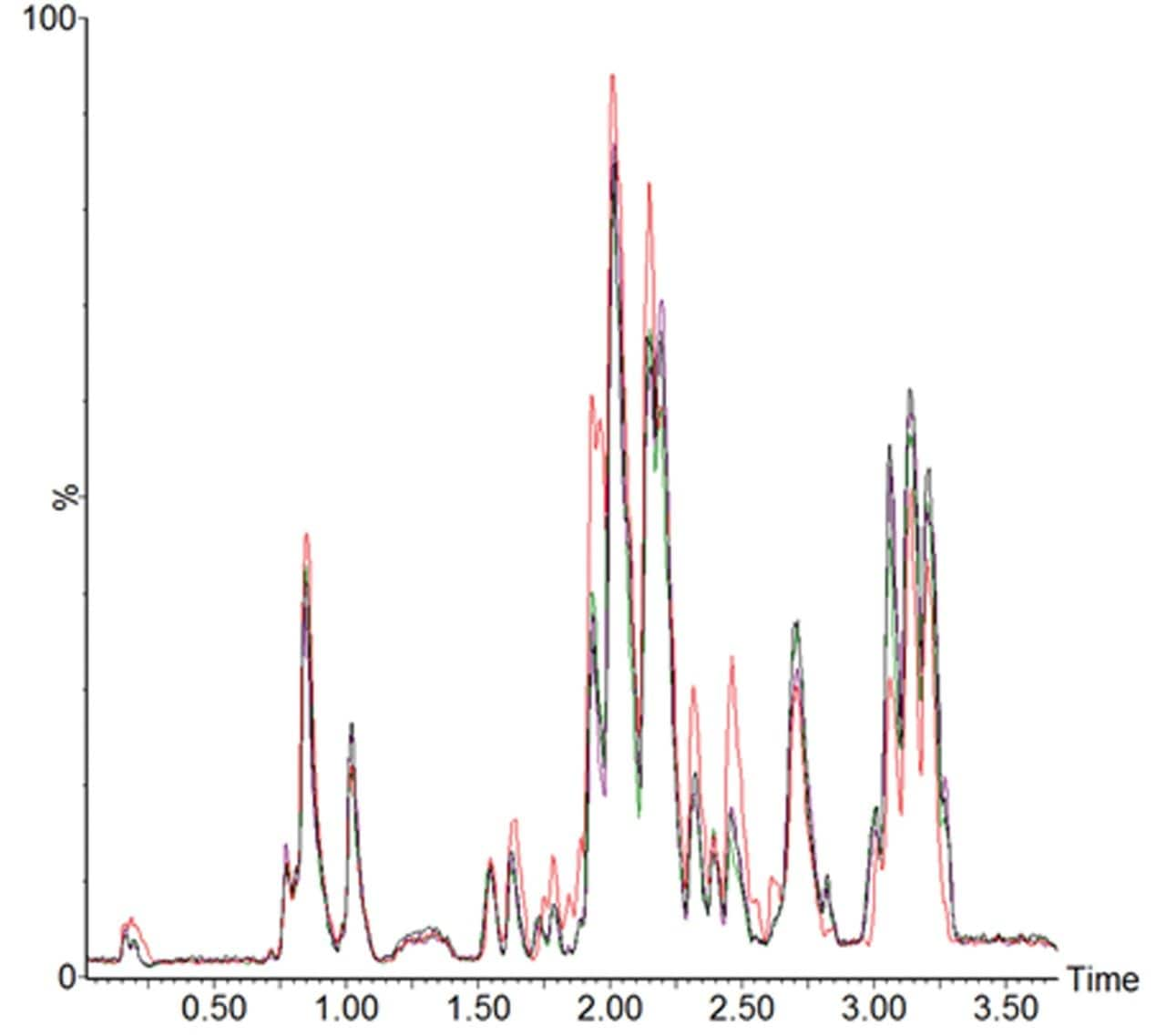
Figure 2 is an example LC-MS chromatogramrepresenting the extracted lipid QC sample,comparing conventional UPLC and RAMMP lipidmethods. Total ion current (TIC) chromatogramsdemonstrate that the chromatographic profileof the lipid analysis was conserved whentransferring from UPLC to RAMMP, maintaininglipid class separation for free fatty acids (FFA),glycerophospholipids and glycerolipids. Thereproducibility of RAMMP was evaluated bycomparing the chromatographic profile of theQC samples throughout the analysis of theplasma batch. The overlaid chromatograms forfour QCs representing the beginning, middle,and end of the analytical batch are displayedin Figure 3. The data demonstrates thereproducibility of the RAMMP Lipid LC-MSmethodology and that the quality of theseparation is not deleteriously affected bythe analysis of multiple plasma samples.
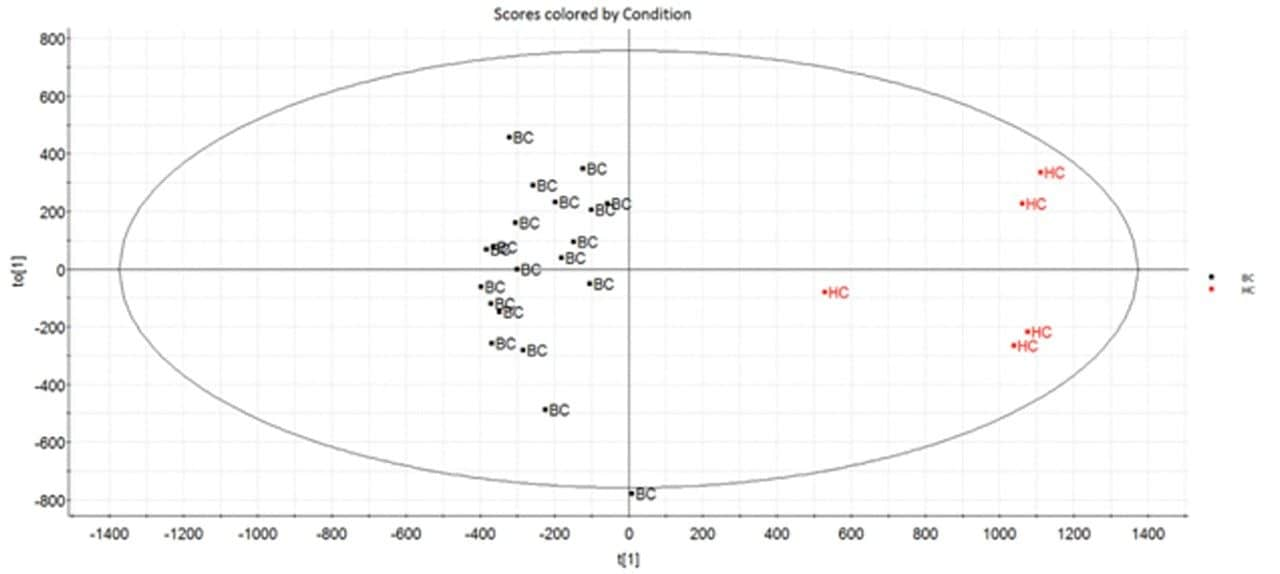
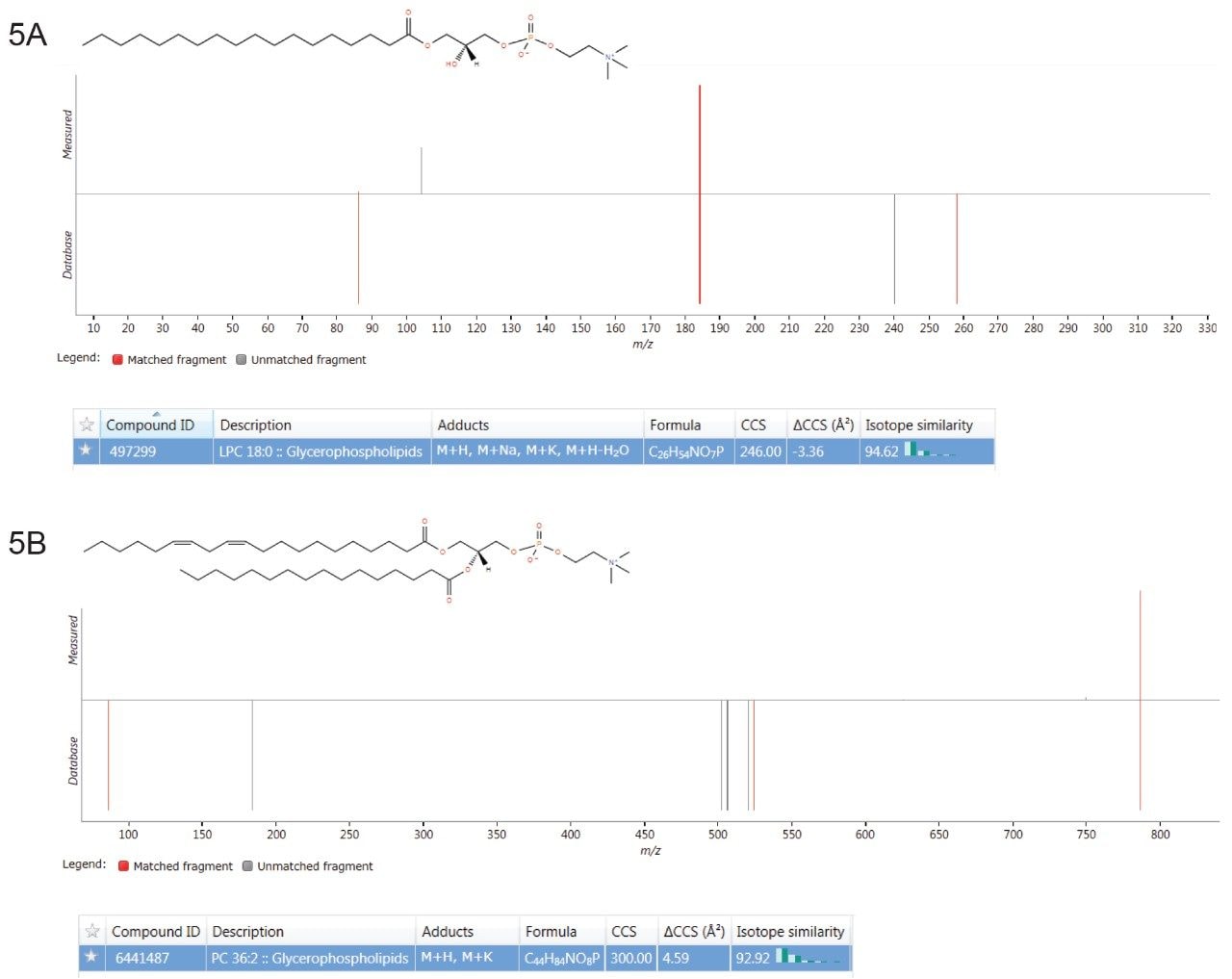
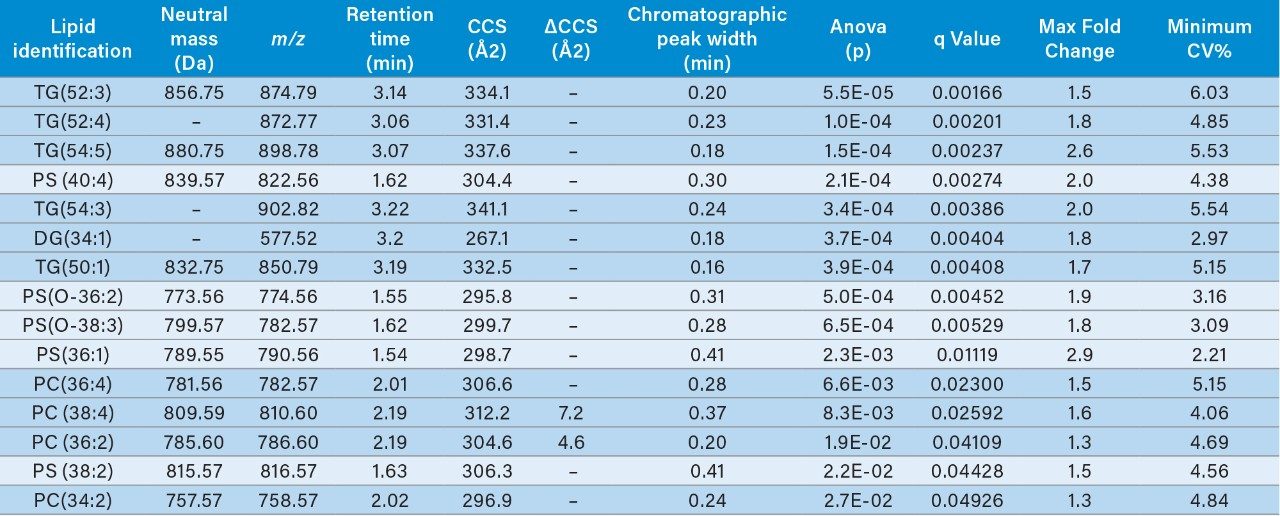
Multivariate statistical analysis (MVA) shows clear separation between breast cancer samples (BC) and healthy controls (HC),when applying an orthogonal partial least squares (OPLS-DA) approach (Figure 4). To ensure that the loss in peak capacitywas minimized when switching from the conventional more extensive UPLC analysis to RAMMP, IMS was implemented aspart of the workflow resulting in increased feature discrimination per unit time. The implementation of IMS improved the spectralclarity and facilitated the measurement of collision cross section (CCS) values. The fragment ion spectra and derived CCSvalues were used to identify differentially expressed lipids within the breast cancer cohort. Features contributing to the MVAdiscrimination were subjected to database analysis. The CCS enabled searches resulted in a number of top hit database searches(Figure 5), including PC 36:2, which was shown to be up regulated in healthy control plasma. The same lipid was also identified asa high ranking hit when searched against Lipidblast and METLIN databases. The lipids detected and shown to be differentiallyexpressed in the breast cancer samples compared to the healthy controls are listed in Table 1. From this data we can see that thephosphatidylcholines lipids, triglycerides, and diglycerides showed lower expression in the breast cancer samples whereas thephosphatidylserine showed greater expression in the breast cancer samples.
1. Kohno S, Keenan AL, Ntambi JM, Miyazaki M.Lipidomic INSIGHT into cardiovascular diseases.Biochem Biophys Res Commun. 2018 Apr 14.
2. Xia J, Wishart DS. Using MetaboAnalyst 3.0 forComprehensive Metabolomics Data Analysis.Curr Protoc Bioinformatics. 2016 55:14.10.1–14.
3. Lewis MR, Pearce JT, Spagou K, Green M,Dona AC, Yuen AH, David M, Berry DJ,Chappell K, Horneffer-van der Sluis V, Shaw R,Lovestone S, Elliott P, Shockcor J, Lindon JC,Cloarec O, Takats Z, Holmes E, Nicholson JK.Development and Application of Ultra-PerformanceLiquid Chromatography-TOF MS for PrecisionLarge Scale Urinary Metabolic Phenotyping.Anal Chem. 2016 88(18):9004–13.
4. Gray N, Adesina-Georgiadis K,Chekmeneva E, Plumb RS, Wilson ID,Nicholson JK. Development of a Rapid MicroboreMetabolic Profiling Ultraperformance LiquidChromatography-Mass Spectrometry Approachfor High-Throughput Phenotyping Studies.Anal Chem.2016;88(11):5742–5.
720006321, June 2018